1. Choose a theme;
2. Click on the corresponding plot;
3. Click on "add";
4. Use the mouse for interactivity and/or edit the code.
Reminder: the editor uses infinite undo/redo in the code (Ctrl+Z / Shift+Ctrl+Z) !Interactive Simulations with JSPlotly
General instructions
1 Simulations
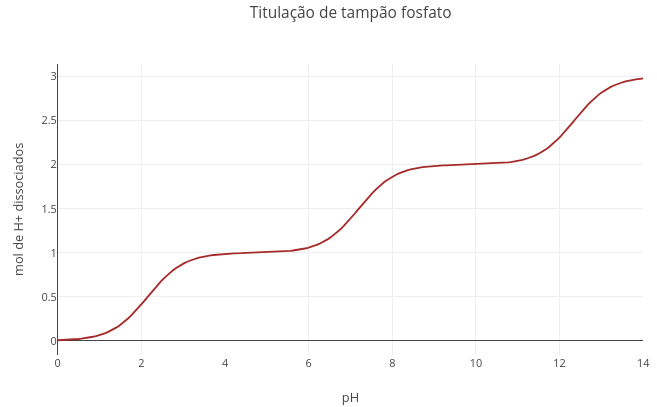
1.1 Acid–base equilibrium and buffer system
Context:
Equation:
\[
fa= \frac{1}{1 + 10^{pKa1 - pH}} + \frac{1}{1 + 10^{pKa2 - pH}} + \frac{1}{1 + 10^{pKa3 - pH}}
\] Where, fa = acid fraction (protonable groups)
Suggestion:
"A. Converting the phosphate buffer curve (triprotic) to bicarbonate buffer (diprotic)"
1. Change the pKa values to those of the bicarbonate buffer: pKa1 = 6.1, and pKa2 = 10.3;
2. Set a very large value for pKa3 (e.g., 1e20).
3. Click on "add plot".
Explanation: pKa is a term that represents the logarithm of a dissociation constant (-log[Ka]). With an extreme value, the denominator also becomes immense, nullifying the term that carries pKa3. In JavaScript and other languages, "e" represents the notation for power of 10.
"B. Converting the bicarbonate buffer curve to acetate"
1. Just repeat the procedure above, with pKa1 = 4.75, and eliminating pKa2.
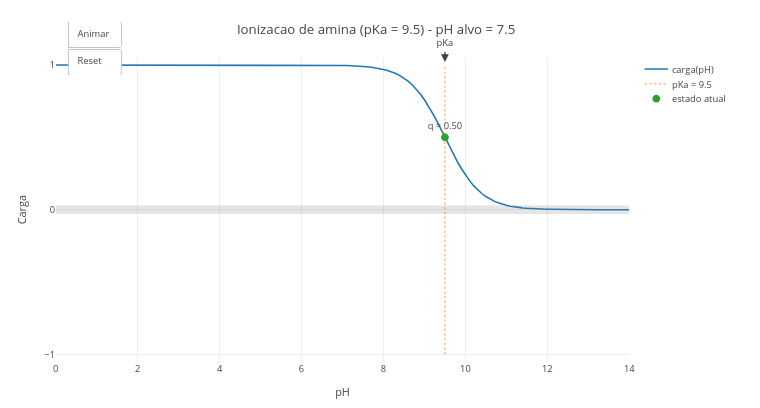
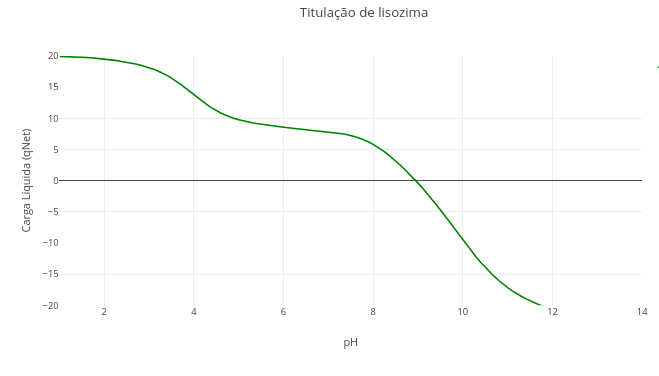
1.2 Ionization state of chemical groups of biomolecules
Context:
Instruction
After clicking the corresponding image below, click on “add” and observe the generated acid–base titration plot. The abscissa axis (x-axis) has an extensive range of pH values, and the ordinate axis the charge that the molecule with ionization potential manifests at each pH, and as a function of its pKa value;
Click on “Animate” to observe the displacement of a small ball along the titration curve, up to a pH value pre-defined in the simulator script, and as a function of its pKa;
Modify in the script the type of ionizable group (in “let grupo_edit =”) and/or the desired final pH (“let pH_final_edit =”), to evaluate the ionization state of a new group;
Click on “Reset” and “Animate”, to observe the new animation.
Suggestion:
1. Change the pH and the type of group that is intended to be observed;
2. Simulate the ionization state of different molecules, such as drugs with carboxylic acid (e.g., acetylsalicylic acid... aspirin), at blood pH and in the gastric cavity;
3. Create a new ionizable group and predict its ionization state. For this, just change the information in 3 constants at the beginning of the script: groups, pKas, and types.
a.
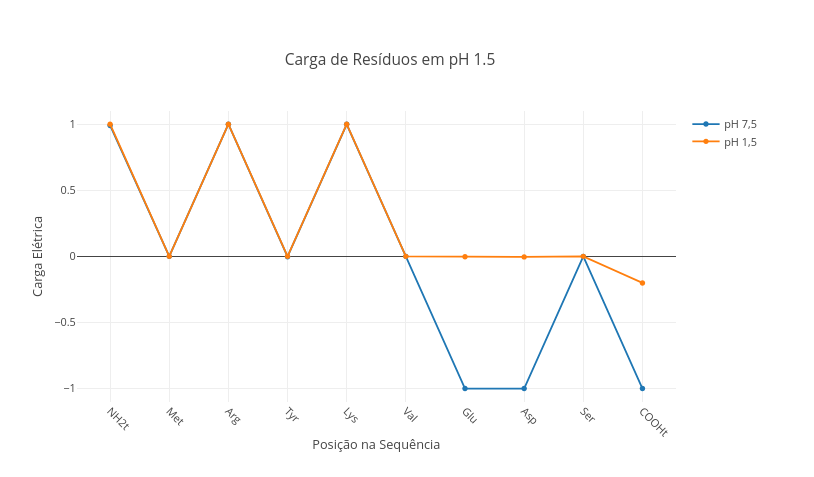
1.3 Net charge network in peptides
Context:
Equation:
\[
q =
\begin{cases}
-\dfrac{1}{1 + 10^{pK_a - pH}} & \text{(acid group)} \\
\dfrac{1}{1 + 10^{pH - pK_a}} & \text{(basic group)}
\end{cases}
\]
Where,
- pKa = value of the base-10 antilogarithm for the equilibrium constant of acid dissociation, Ka (or log[Ka]).
Suggestion:
1. Select the peptide sequence below, and observe the charge distribution:
"Ala,Lys,Arg,Leu,Phe,Glu,Cys,Asp,His"
2. Simulate the stomach pH condition ("const pH = 1.5"), and verify the change of charges in the peptide.
3. Select a physiological peptide (oxytocin, for example), observe its charge in blood (pH 7.5), and reflect on its potential for electrostatic interaction with cellular components.
"Cys,Tyr,Ile,Gln,Asn,Cys,Pro,Leu,Gly" - oxytocin
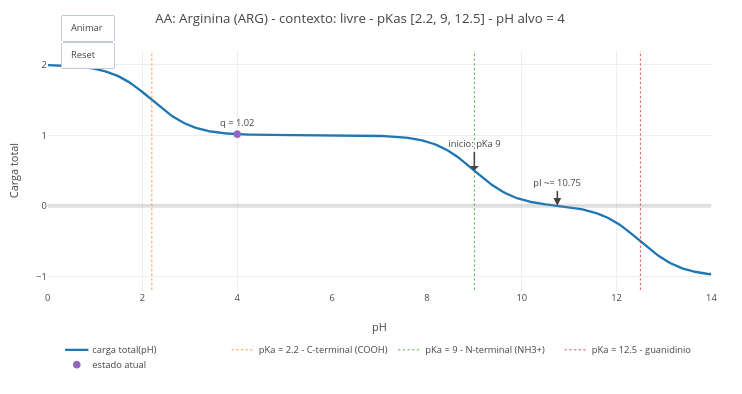
1.4 Charge, pH, pKa & pI of amino acids (animation)
Context:
Suggestion:
1. Simulate the condition of an amino acid at blood pH and at stomach pH;
2. Compare the pH values of a free amino acid, with that of its residue in a chain;
3. Change the boolean option in "const showSites", to observe the curves of each ionizable group.
1.5 Isoelectric Point in Proteins
Context:
Equation:
\[ q_{\text{net}}(pH) = \sum_{i=1}^{N} \left[ n_i \cdot q_{B_i} + \frac{n_i}{1 + 10^{pH - pK_{a_i}}} \right] \]
Where,
- qnet = total net charge;
- qB$_{i} = charge of the basic form for residue i (for example, +1 for Lys, 0 for Asp);
- n\(_{i}\) = number of groups of residue i.
Suggestion:
"Finding the pI for other proteins"
1. One can verify the titration of any other protein or peptide sequence by simply substituting the primary sequence contained in the code. An assertive way to perform this substitution involves:
a. Search for the "FASTA" sequence of the protein in NCBI ("https://www.ncbi.nlm.nih.gov/protein/") - e.g.: "papain";
b. Click on "FASTA" and copy the sequence obtained in 1a.;
c. Paste the sequence into a site for residue quantification (e.g., "https://www.protpi.ch/Calculator/ProteinTool");
4. Substitute the sequence in the code.
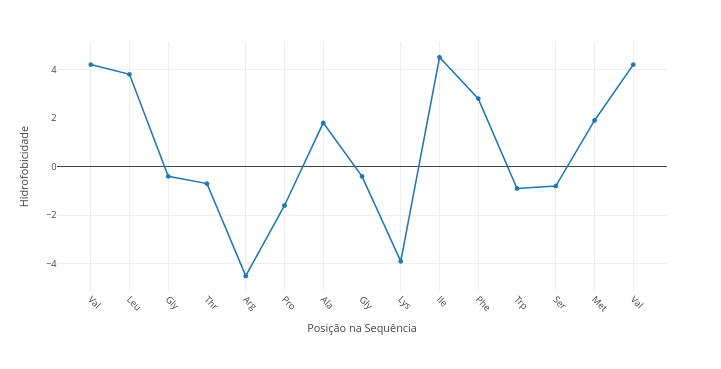
1.6 Hydrophobicity in amino acid sequences
Context:
Equation:
\[
\tilde{y}*i ;=; \frac{1}{w} \sum*{k=i-m}^{i+m} y_k,
\qquad w = 2m+1
\]
Where,
\[\begin{aligned} y_i &:; \text{hydrophobicity value at position $i$ of the sequence} \ \tilde{y}_i &:; \text{smoothed value (moving average) at position $i$} \ m &:; \text{half-window (number of neighbors on each side)} \ w &:; \text{total window width ($w=2m+1$)} \ i &:; \text{amino acid position in the sequence ($1 \leq i \leq N$)} \end{aligned}\]:;
m &:;
w &:;
i &:; \end{aligned}
Suggestion:
1. Try changing the sequence to a known one;
2. Compare a polar sequence with a nonpolar one by overlay ("add");
3. Transcribe a transmembrane helix sequence of a protein by accessing its data on the PDB site
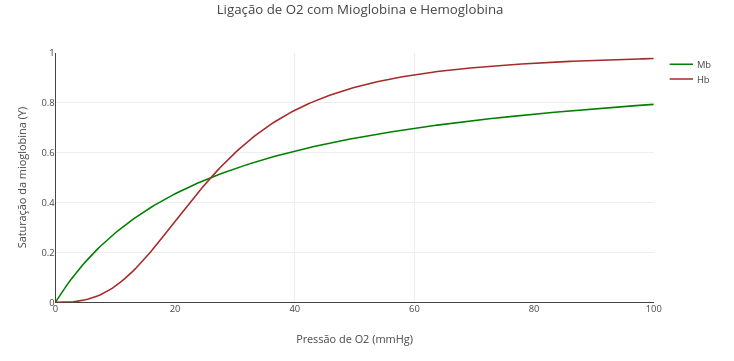
1.7 Interaction of oxygen with myoglobin and hemoglobin
Context:
Equation:
\[
Y= \frac{pO_2^{nH}}{p_{50}^{nH} + pO_2^{nH}}
\]
Where
- Y = degree of oxygen saturation in the protein;
- pO\(_{2}\) = oxygen pressure;
- p\(_{50}\) = oxygen pressure at 50% saturation;
- nH = Hill coefficient for the interaction;
Suggestion:
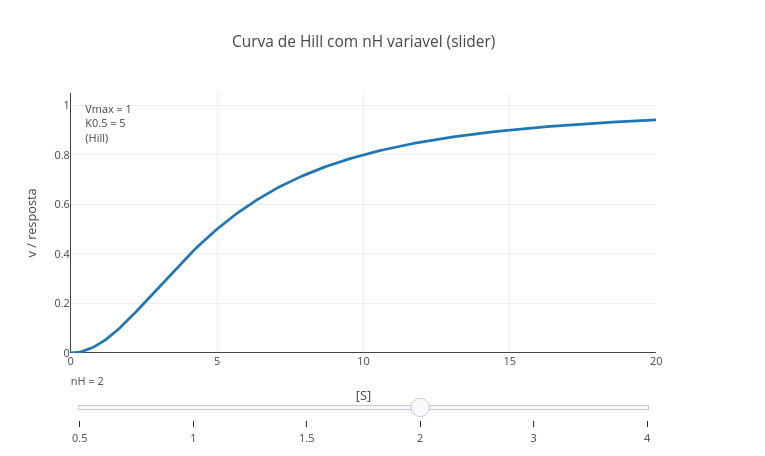
1. Run the application ("add plot"). See that the value of "nH" of the Hill constant is "1", that is, without cooperativity effect.
2. Now replace the value of "nH" with the Hill coefficient for hemoglobin, 2.8, and run again !
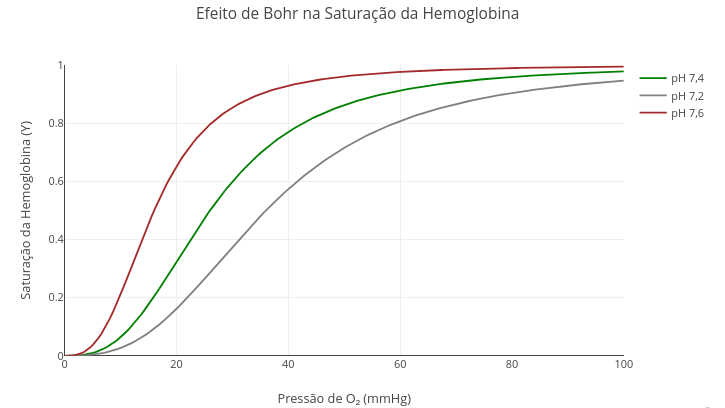
1.8 Bohr effect in hemoglobin (pH)
Context:
Equation:
\[ Y(pO_2) = \frac{{pO_2^n}}{{P_{50}^n + pO_2^n}}, \quad \text{with } P_{50} = P_{50,\text{ref}} + \alpha (pH_{\text{ref}} - pH) \]
Where,
- Y = hemoglobin saturation,
- pO\(_{2}\) = partial pressure of oxygen (in mmHg),
- P\(*{50}\) = pressure of O\(*{2}\) at which hemoglobin is 50% saturated,
- P\(_{50,ref}\) = 26 mmHg (standard value),
- \(\alpha\) = 50 (intensity of the Bohr effect),
- pH\(_{ref}\) = 7.4,
- n = 2.8 = Hill coefficient for hemoglobin.
Suggestion
1. Try altering the reference pH for the interaction;
2. Simulate other allosteric models by changing the value of "n"
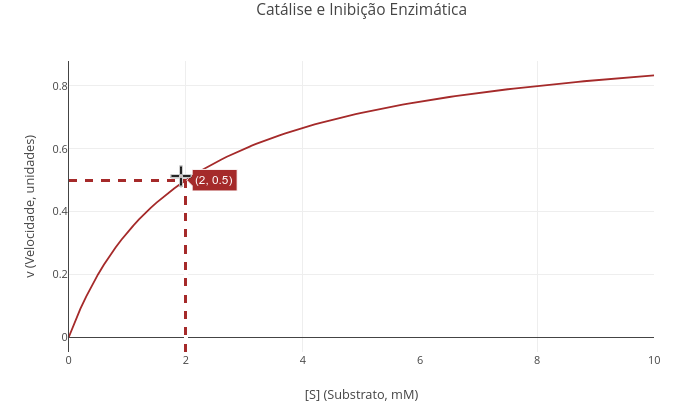
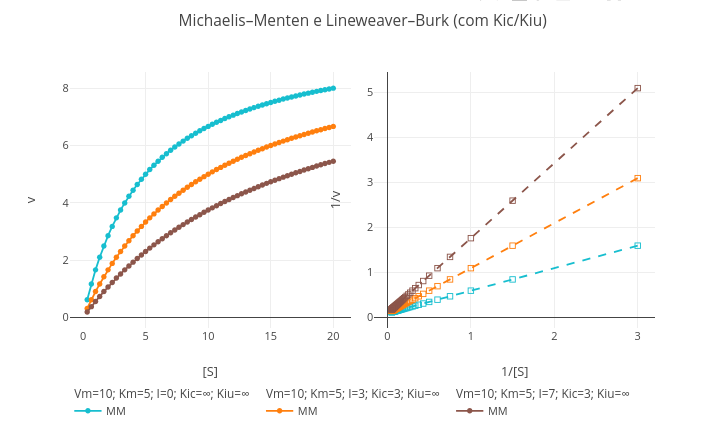
1.9 Catalysis and enzymatic inhibition
Context:
Equation:
\[ v=\frac{Vm*S}{Km(1+\frac{I}{Kic})+S(1+\frac{I}{Kiu})} \]
Where
- S = substrate content for the reaction;
- Vm = limiting velocity of the reaction (in books, maximum velocity);
- Km = Michaelis–Menten constant;
- Kic = inhibitor dissociation equilibrium constant for the competitive model;
- Kiu = inhibitor dissociation equilibrium constant for the uncompetitive model
Suggestion:
"A. Enzymatic catalysis in the absence of inhibitor."
1. Just run the application with the general equation. See that the values for Kic and Kiu are high (1e20). In this way, with high "dissociation constants", the interaction of the inhibitor with the enzyme is irrelevant, returning the model to the classic Michaelis–Menten equation.
2. Try changing the values of Vm and Km, comparing plots.
3. Use the geographic coordinates feature of the icon bar ("Toggle Spike Lines"), to consolidate the mathematical meaning of Km, as well as observe the effect of distinct values of this on the visualization of the plot.
"B. Competitive inhibition model."
1. To observe or compare the michaelian model with the competitive inhibition model, just replace the value of Kic with a consistent number (e.g., Kic= 3).
"C. Uncompetitive inhibition model."
1. The same suggestion above serves for the uncompetitive model, this time replacing the value for Kiu.
"D. Pure noncompetitive inhibition model."
1. In this model, the simulation is given by equal values for Kic and Kiu.
"E. Mixed noncompetitive inhibition model."
1. For this model, just allocate distinct values for Kic and Kiu.
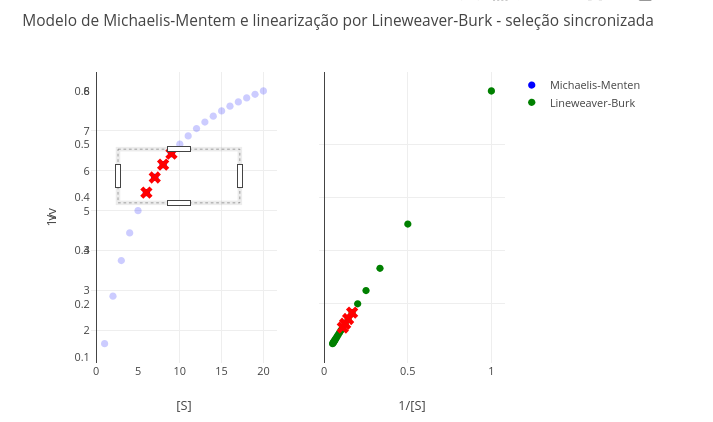
1.10 Linearization of kinetic data by Lineweaver–Burk
Context
1.10.1 Equation
\[
\frac{1}{v} = \frac{1}{S}*\frac{Km}{Vm} + \frac{1}{Vm}
\]
Suggestion
1. Try selecting the extremes of the Michaelis–Menten plot (hyperbolic curve), and observe the same points in the double-reciprocal. Which values are more reliable in the latter, the first or the last?
2. Research other forms of linearization (e.g., Eadie–Hofstee; "v/S" X "v"), and see how the transformation and point selection presents itself. For this, change the constants below:
const invS = S.map(s => 1/s);
const invV = V.map(v => 1/v);
1.11 Diagnosis of Enzymatic Inhibition (Michaelis–Menten and Lineweaver–Burk)
Context
Suggestion
1. Simulate the conditions for a competitive inhibition, inserting a value for the inhibitor content ("I") and changing the value of "Km";
2. Do the same for an uncompetitive model, although changing the value of "Vm";
3. Try the pure noncompetitive model, changing "Km" and "Vm" to the same value;
4. Test mixed competitive inhibition, inserting distinct values for "Km" and for "Vm."
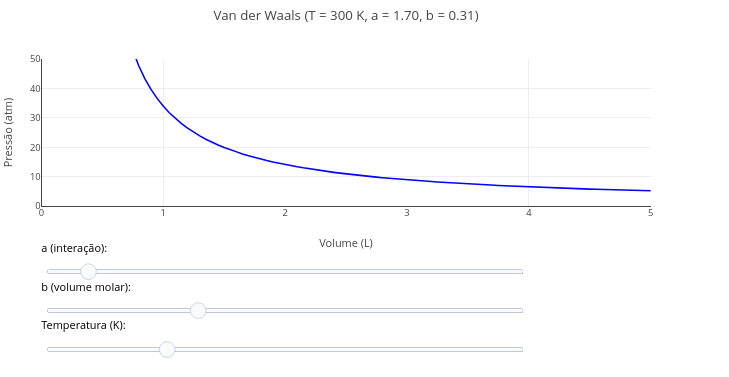
1.12 van der Waals equation for ideal gases
Context:
Equation:
\[ P = \frac{RT}{V - b} - \frac{a}{V^2} \]
- P = gas pressure (atm);
- V = molar volume (L);
- T = temperature (K);
- R = 0.0821 = ideal gas constant (L·atm/mol·K);
- a = intermolecular attraction constant (L\(^{2}\)·atm/mol$^{2})
- b = excluded volume constant (L/mol)
Suggestion:
1. Try varying the parameters of the equation by means of the "slider" for temperature, as well as for the coefficients "a and b".
2. Discuss which of the coefficients has the greatest effect on the curve profile, and the reason for this.
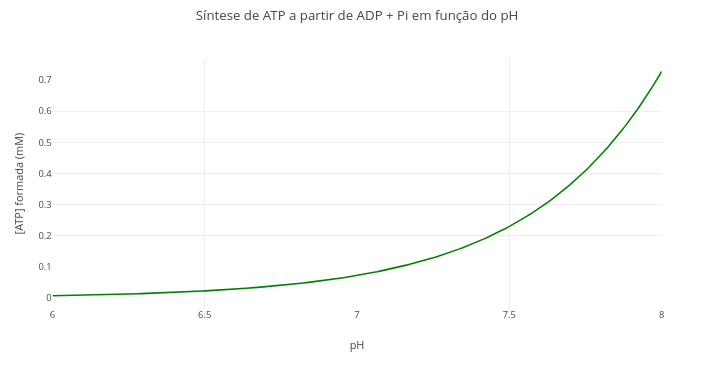
1.13 Equilibrium of ATP production from reagents, temperature, and pH
Context:
Equation:
\[ \Delta G = \Delta G^{\circ'} + RT \ln\left(\frac{[\text{ADP}] \cdot [\text{P}_i]}{[\text{ATP}]}\right) + 2{,}303 \cdot RT \cdot n_H \cdot \text{pH} \]
Where,
- \(\Delta\)G = Gibbs energy of the reaction (positive for spontaneously unfavorable synthesis, kJ/mol);
- \(\Delta\)G\(^{o'}\) = 30.5 kJ/mol biological standard Gibbs energy for ATP synthesis;
- R = 8.314 J/mol/K (general gas constant);
- T=310 K (physiological temperature);
- nH\(^{+}\) = 1 (number of protons involved in the reaction);
- [ADP], [Pi], [ATP] = molar concentrations of reagents and product
Suggestion:
1. Change the quantities involved in the expression, and compare with previous visualizations. For example, temperature, pH, and ADP and Pi contents.1.14 Variation of Gibbs energy with temperature
Context:
Equation:
\[ \Delta G(T) = \Delta H^\circ - T,\Delta S^\circ + \Delta C_p \left(T - T_0 - T \ln\left(\frac{T}{T_0}\right)\right) \]
Where,
- \(\Delta\)G(T) = Gibbs energy of the reaction at each temperature value, kJ/mol);
- \(\Delta\)H\(^{o}\) = standard enthalpy of the reaction at T\(_{0}\), usually 298 K (J/mol);
- \(\Delta\)S\(^{o}\) = standard entropy of the reaction at T\(_{0}\);
- \(\Delta\)Cp = change in heat capacity of the reaction (J/mol·K), assumed constant with temperature;
- T = temperature of interest (K);
- T\(_{0}\) = reference temperature, generally 298 K.
- R = 8.314 J/mol/K (general gas constant);
Suggestion:
1. Try varying one or more parameters of the expression;
2. Test the behavior of the Gibbs curve at an elevated reference temperature (simulation for an extremophile organism);
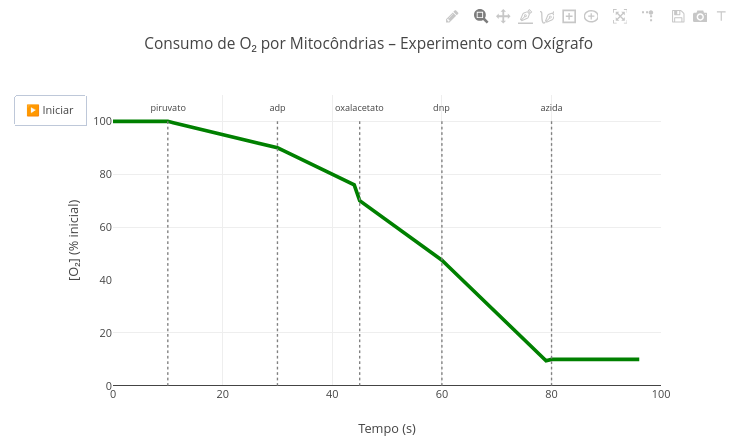
3. Simulate the situation where the heat capacity change is null1.15 Oxygen consumption by mitochondria
Context:
Suggestion:
1. Note that the appearance of metabolites and the resulting oxygen consumption rates are defined by the function "oxigenio". Try changing them and observe the effect in the animation;
2. Realize that the metabolites and their action, as well as the measured signal, O2 content, are easily adapted for any other metabolic measurement in the code. Try, for example, what the plot would look like in a simulation for the glycolytic pathway;
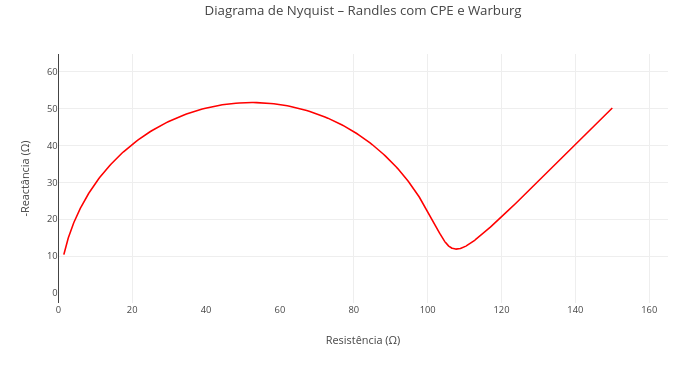
3. This code was designed for an animation. Go to the "Applications" section below, and try "FlowForces" for a similar purpose, although with full interactive adaptation.1.16 Electrochemical impedance spectroscopy (EIS)
Context:
Equation:
\[ Z_{\text{total}}(\omega) = R_s + \left[ \left( \frac{1}{R_p} + Q (j\omega)^n \right)^{-1} \right] + \frac{\sigma}{\sqrt{\omega}} (1 - j) \]
Where:
- Z\(_{total}\) = total impedance at a given frequency ;
- \(\omega\) = angular frequency (rad/s), \(\omega\)=2πf or \(\omega\)=2πf;
- R\(_{p}\) = ohmic resistance (resistance of the electrolyte solution, wires, contacts etc.) ;
- R\(_{p}\) = polarization resistance (associated with charge-transfer processes, such as electrochemical reactions);
- Q = constant associated with the constant phase element (CPE), replaces an ideal capacitor to represent non-ideal behaviors ;
- n = CPE exponent, between 0 and 1; defines the degree of ideality of the capacitive behavior (n = 1: ideal capacitor; n < 1: dispersion);
- \(\sigma\) = Warburg coefficient, associated with ion diffusion in the electrochemical system ;
- j = imaginary unit, j\(^{2}\) = −1;
Suggestion
1. Check the effect of Rs on the plot, nullifying its value (solution resistance);
2. Observe the deformation of the semicircle by varying the values of the constant phase element (e.g., Q = 1e-3; n = 0.6 - dispersion of the capacitive behavior);
2. Try combining other values of the parameters in the code header, to highlight common situations in electroanalysis: Rs, Rp, Q, n, and sigma;
3. Reduce the Warburg-with-constant-phase-element model to a simple Randles model, formed only by 2 resistors in series, the second in parallel with an ideal capacitor, and without Warburg diffusion (sigma = 0; n = 1).1.17 Cyclic voltammetry
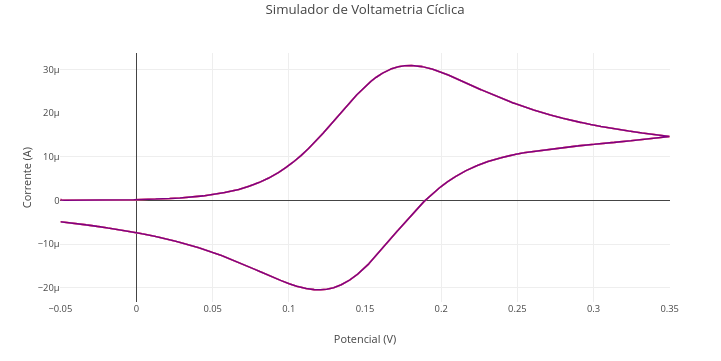
Context:
Equation:
\[ j = j_0 \left[ \exp\left(\frac{\alpha n F (E - E^0)}{RT}\right) - \exp\left(\frac{-(1 - \alpha) n F (E - E^0)}{RT}\right) \right] \] Where:
- j = current density;
- j\(_{0}\) = exchange current;
- \(\alpha\) = charge transfer coefficient;
- E = potential applied to the electrode;
- E\(^{0}\) = standard electrode potential;
- n = number of electrons;
- F = Faraday constant (96485 C·mol⁻¹)
- R = general gas constant (8.314 J·mol⁻¹·K⁻¹);
- T = temperature
\[ \frac{\partial C(x,t)}{\partial t} = D \frac{\partial^2 C(x,t)}{\partial x^2} \]
Where:
- C(x,t) = concentration of the electroanalytical species (mol/cm³), as a function of position x and time t;
- D = diffusion coefficient of the species (cm²/s);
- t = time (s);
- x = distance from the electrode surface (cm);
- \(\delta\) = notation for partial derivative.
\[ E = E^0 + \frac{RT}{nF} \ln \left( \frac{[\text{Ox}]}{[\text{Red}]} \right) \]
\[ i(t) = n F A D \left. \frac{\partial C(x,t)}{\partial x} \right|_{x=0} \] Where:
- A = electrode surface area (cm²);
- i(t) = current at time t (A, Ampere).
Cyclic Voltammetry Simulator
Suggestion:
1. Observe the number of values at the beginning of the code, tangible to a "parametric manipulation". Try to know what they represent, and try to vary their values consciously, aiming to add value to learning the simulation. This is the soul of "parametric manipulation" that involves "reproducible teaching" !!1.18 Diagrams and flowcharts
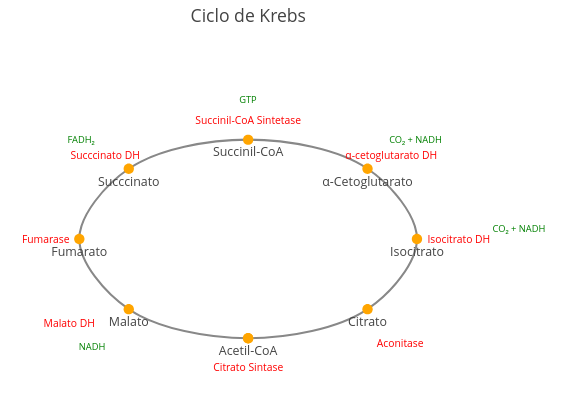
Context: Diagram
Suggestion:
1. Try repositioning enzymes and metabolites better, just by clicking and dragging the terms;
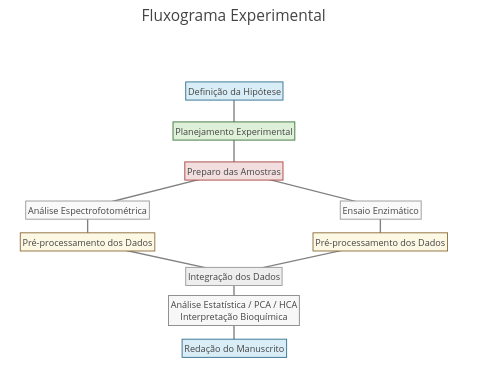
2. Try replacing the names that are in the code to produce another metabolic cycle, such as the urea cycle.Context: Flowchart
Suggestion:
1. Reposition terms and connectors by mouse dragging;
2. For a flowchart different in content, modify the terms in the code;
2. For a flowchart different in format, change the font and connector characteristics in the "annotations" constant.
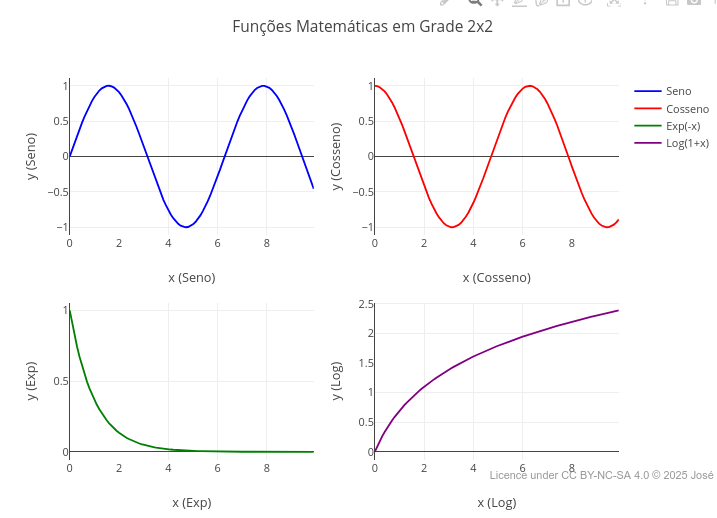
2 Construction of plots
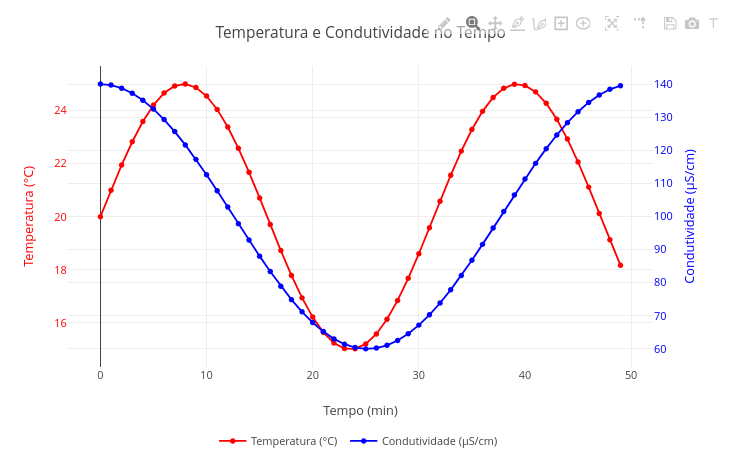
Plotly.js library is prodigious in elaborating a significant set of varied types, as represented in its version for the R language on the developer site.2.1 Plot with 2 y-axes
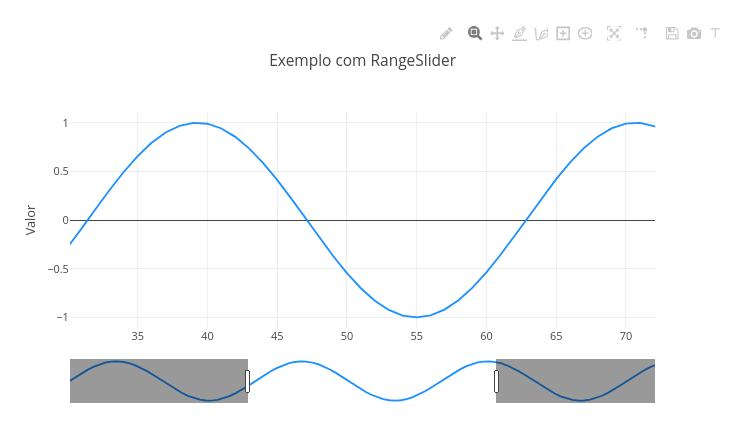
2.2 RangeSlider
2.3 Subplots
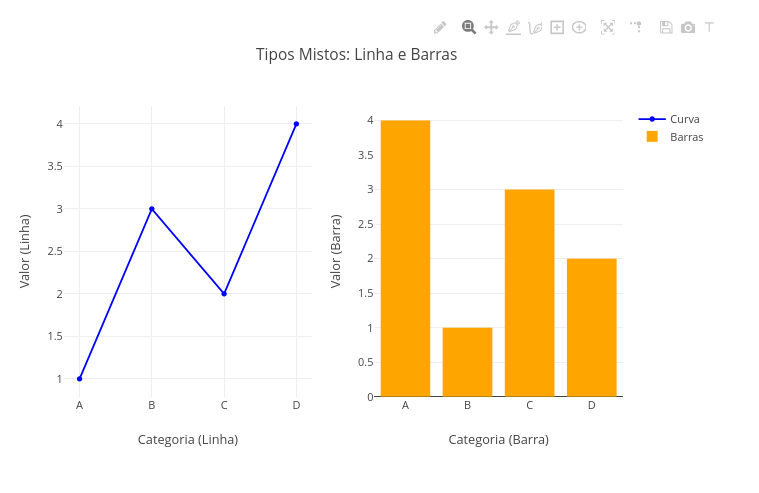
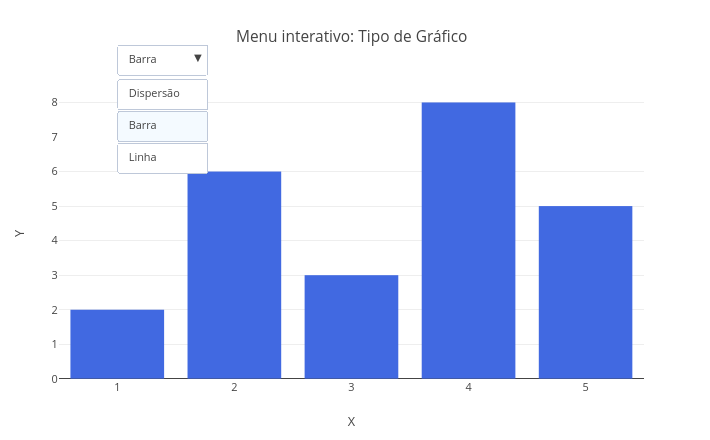
2.4 Subplots with different types
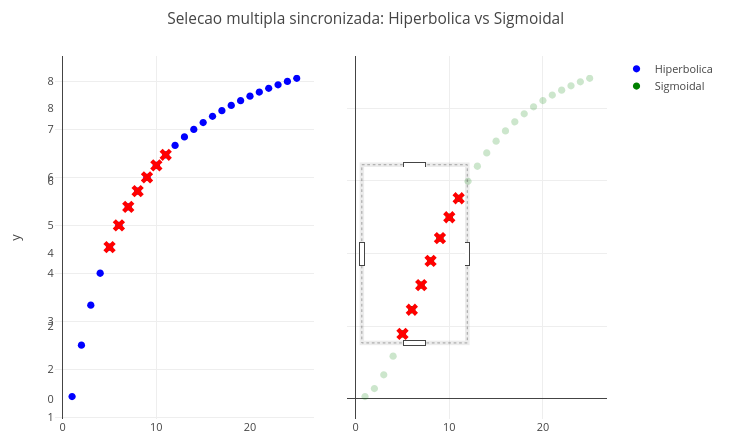
2.5 Synchronized multiple selection
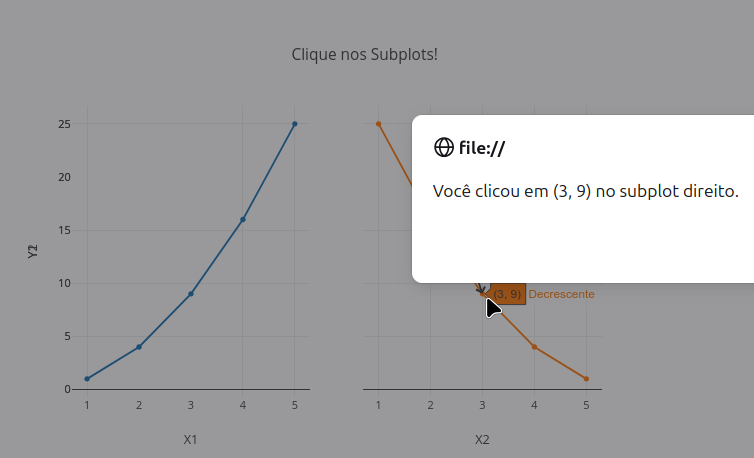
2.6 Click event & message
2.8 Slider control
3 Data analysis
3.1 User data entry
Context
Suggestion:
1. Try changing the inserted data, overlaying the plot or not;
2. Try changing the data representation in "mode" and "type", to points, lines, points+lines, bars.
3.2 Loading a file for analysis
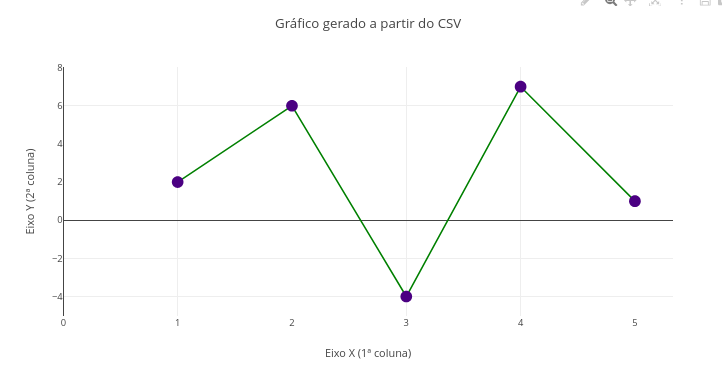
Context - CSV file
Instructions:
- 1 Click add plot and select a CSV file using the upper browse button that is shown. Note: variable X in the 1st column of the file, and variable Y in the 2nd column;
- Click add plot again to view the resulting plot.
Suggestion
1. Try other CSV files;
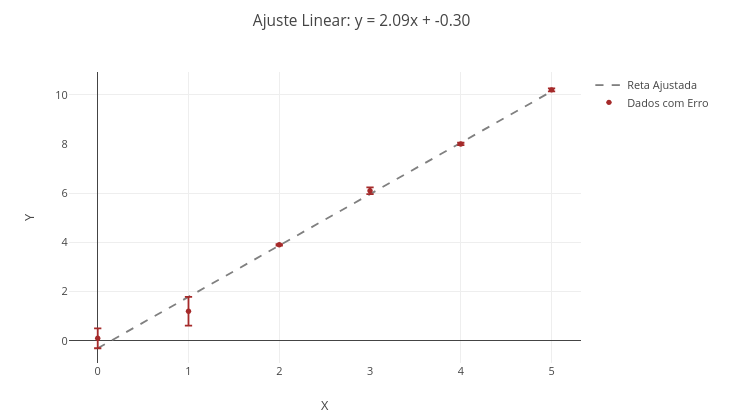
2. Vary aspects of the plot, such as type, color, marker size, etc.3.3 Linear fitting of data
Context:
Equation:
\[ y = \alpha x + \beta + \varepsilon \]
Where:
- y = dependent variable;
- x = independent variable;
- \(\alpha\) = slope of the fitted line (slope);
- \(\beta\) = intercept of the fitted line;
- \(\epsilon\) = measurement error.
\[ \alpha = \frac{n \sum x_i y_i - \sum x_i \sum y_i}{n \sum x_i^2 - \left( \sum x_i \right)^2} \]
\[ \beta = \frac{\sum y_i - \alpha \sum x_i}{n} \]
\[ \hat{y}_i = \alpha x_i + \beta \]
\[ \varepsilon_i = \left| y_i - \hat{y}_i \right| \]
Suggestion:
1. Show the points overlaid or not with the fitted line. For overlay, choose "mostrarPontos = true" and "mostrarReta = true";
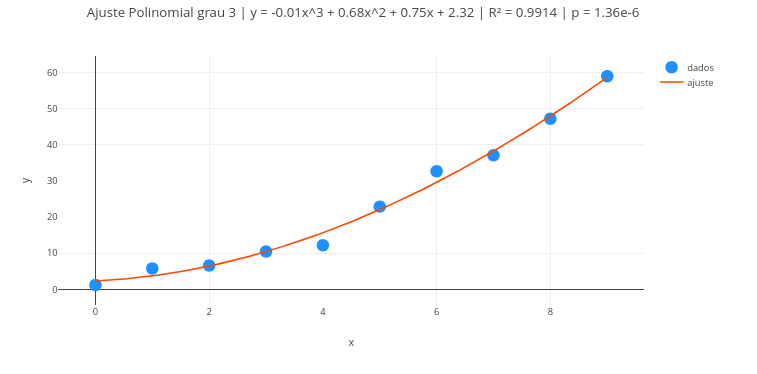
2. Change the data and perform a new fit, to obtain other line parameters.3.4 Polynomial regression
Context:
3.4.1 Equation
\[ y = \beta_0 + \beta_1 x + \beta_2 x^2 + \cdots + \beta_g x^g \] | Thus, the Vandermonde matrix is built as:
\[ X = \begin{bmatrix} 1 & x_1 & x_1^2 & \cdots & x_1^g \\ 1 & x_2 & x_2^2 & \cdots & x_2^g \\ \vdots & \vdots & \vdots & & \vdots \\ 1 & x_n & x_n^2 & \cdots & x_n^g \end{bmatrix} \]
\[
\boldsymbol{\beta} = (X^T X)^{-1} X^T \mathbf{y}
\]
Where:
- T represents the transpose matrix
Suggestion:
1. Try degree 1 for the polynomial, i.e., reduce the treatment to a linear fit;
2. Change the formatting of labels, colors, sizes, etc., in the code;
3. Overlay some fits, edit and reposition the legend;
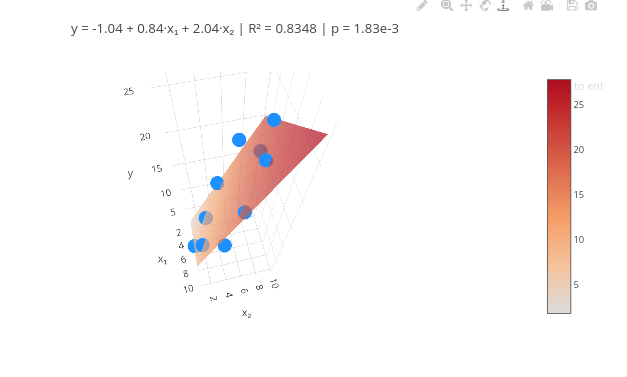
4. Test the code with another data vector.3.5 Multiple linear regression
Context:
Equation:
\[ y_i = \beta_0 + \beta_1 x_{1i} + \beta_2 x_{2i} + \cdots + \beta_p x_{pi} + \varepsilon_i \]
\[ \mathbf{y} = \mathbf{X} \boldsymbol{\beta} + \boldsymbol{\varepsilon} \]
\[ \hat{\boldsymbol{\beta}} = (\mathbf{X}^\top \mathbf{X})^{-1} \mathbf{X}^\top \mathbf{y} \]
Where:
- \(\beta\) = vector of coefficients;
- y = response vector;
- X = design matrix;
- \(\epsilon\) = random noise
\[ \hat{\mathbf{y}} = \mathbf{X} \hat{\boldsymbol{\beta}} \] \[ \mathbf{e} = \mathbf{y} - \hat{\mathbf{y}} \]
Instructions
Note that there is a boolean flag (mostrarAjuste) at the beginning of the code: false for data only, and true for the fit;
You can click add plot to view the data with the flag set to false, followed by another add plot with the flag set to true.
Suggestion:
1. QSAR ("Quantitative Structure-Activity Relationship") results use multiple linear fitting analysis to identify the strength of predictor variables. Search online for a dataset that has 2 variables (e.g., concentration, pH, compound A, B, etc.)
2. For more predictor variables, see the code that follows !!3.6 Multiple linear fitting with 3 or more predictor variables
Context
Suggestion:
1. Try varying the number of predictors (xi). To do this:
a. If you want to reduce, just remove the desired vector(s), and correct their quantity in the line: "const X = x1.map((_, i)";
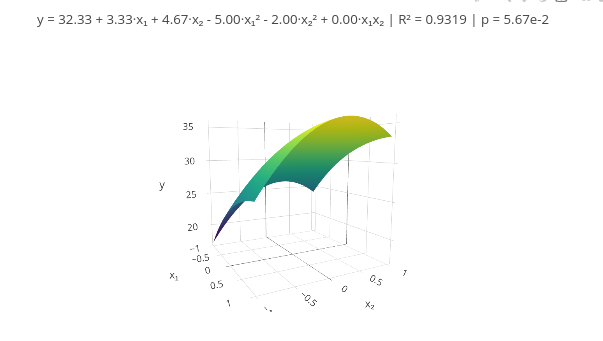
b. If you want to increase, add a new vector in the corresponding line, and update the quantity in the mapping of "const X = x1.map((_, i)".3.7 Response Surface Methodology (RSM)
Context
Equation:
\[ y = \beta_0 + \beta_1 x_1 + \beta_2 x_2 + \beta_{11} x_1^2 + \beta_{22} x_2^2 + \beta_{12} x_1 x_2 + \varepsilon \]
\[ \mathbf{y} = \mathbf{X} \boldsymbol{\beta} + \boldsymbol{\varepsilon} \]
\[
\hat{\boldsymbol{\beta}} = (\mathbf{X}^\top \mathbf{X})^{-1} \mathbf{X}^\top \mathbf{y}
\]
Suggestion:
1. As suggested earlier, try using data from the literature or another source whose responses are already known, replacing the respective vectors at the beginning of the code. This allows comparing the effectiveness of using the presented tool.3.8 Data smoothing - Spline and Savitzky-Golay filter
Context - Cubic spline
Equation
Mathematically, the spline is applied such that:
\[ S_i(x) = a_i + b_i(x - x_i) + c_i(x - x_i)^2 + d_i(x - x_i)^3 \]
Where:
- S\(_{i}\)(x) = cubic spline function in the interval [xi,xi+1];
- a\(*{i}\), b\(*{i}\), c\(*{i}\), d\(*{i}\) = coefficients specific to segment i;
- x\(_{i}\) = starting point of the interval;
- x = independent variable.
Context - Savitzky-Golay filter
Equation
Mathematically:
\[ \tilde{y}*i = \sum*{j=-m}^{m} c_j \cdot y_{i+j} \]
Where:
- y\(_{i}\) = smoothed value at position i;
- y\(_{i+j}\) = real series values within the window;
- c\(_{i}\) = filter coefficients derived from a polynomial regression;
- m = number of points on each side of the central window.
Instructions:
- The code has two true/false flags, one for “usarGolay” and another for “usarSpline”;
- View the raw data adopting false for both flags;
- To overlay a cubic spline, change its constant to true;
- To overlay the Savitzky-Golay filter, change its constant to true;
- To tune the filter, modify its parameters in the constants janela (moving window) and/or grau (polynomial degree).
Suggestion:
1. Compare the smoothing effects of the two polynomial interpolation treatments;
2. Apply only the filter, tuning moving-window and polynomial-degree parameters;
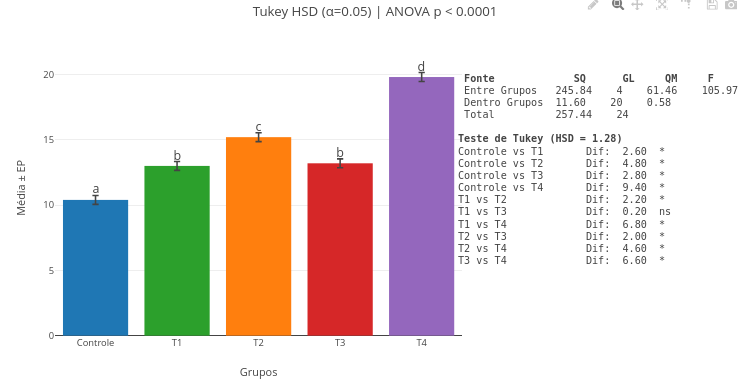
3. Try interpolation with other raw data in the code, replacing the constants "x_values" and "y_raw" with numeric vectors;3.9 ANOVA and Tukey-HSD test
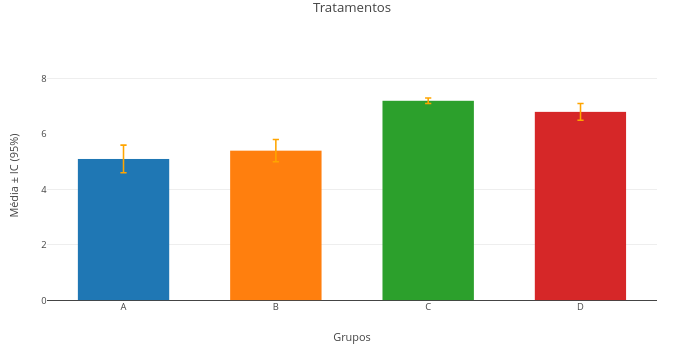
Context:
Equation:
\[ \text{HSD}*{ij} = q*{\alpha} \cdot \sqrt{\frac{\text{MS}_{\text{within}}}{2} \left( \frac{1}{n_i} + \frac{1}{n_j} \right)} \]
Where:
- HSD = minimum value of the difference between two means for it to be considered significant;
- q\(_{\alpha}\) = critical value of the studentized q distribution (Tukey distribution), depending on the number of groups and the significance level \(\alpha\);
- MS\(_{within}\) = mean square within groups (error) obtained from ANOVA;
- n: number of observations per group (i and j if there are groups of different sizes).
Suggestion:
1. You can click and drag the minimum-difference markers (a,b,etc) above the plot for better visual adjustment, as well as the ANOVA table and the statistical test result;
2. Change the vector values, redo the analysis and the plot simultaneously, and notice the difference in the obtained values. Note that, as stated on the lower border, it is necessary to refresh the site page to clear previous data retained in cache memory;
3. Enter other data in the respective vectors, yours or from another source;
4. Insert new data vectors, or delete one, for a customized calculation and plot.3.10 Nonlinear fitting of data
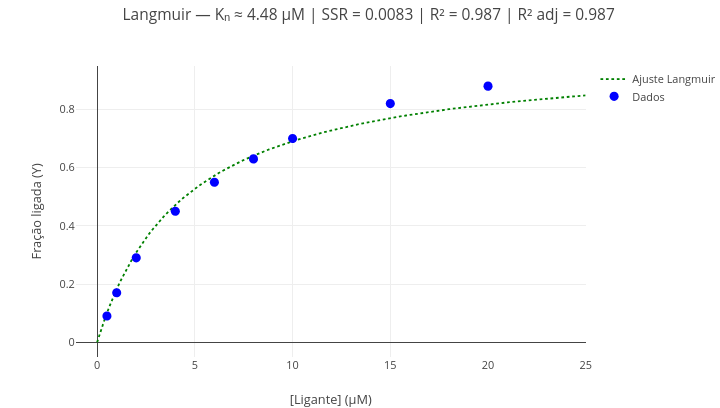
Context: Ligand-protein interaction (grid search algorithm)
Equation:
\[ Y = \frac{aX}{K_d + X} \] Where:
- Y = fraction of occupied sites;
- X = concentration of free ligand;
- a = saturation value (sites of the same affinity fully occupied);
- Kd = equilibrium dissociation constant of the complex.
\[ SSR(a, K_d) = \sum_{i=1}^n \left( y_i - \frac{a x_i}{K_d + x_i} \right)^2 \]
Where:
SSR = sum of squared residuals (inversely proportional to fit quality)
Suggestion:
1. As in the previous example, alternatively show the points and the fitted curve using the boolean operation ("mostrarPontos = true/false"; "mostrarCurva = true/false");
2. Change the data and perform a new fit, to obtain other equation parameters (Kd, SSR, R², adjusted R²). Note: adjusted R² corresponds to the R² value corrected for the number of model parameters (in this case, "2"), directly affecting the "degrees of freedom" for the fit.
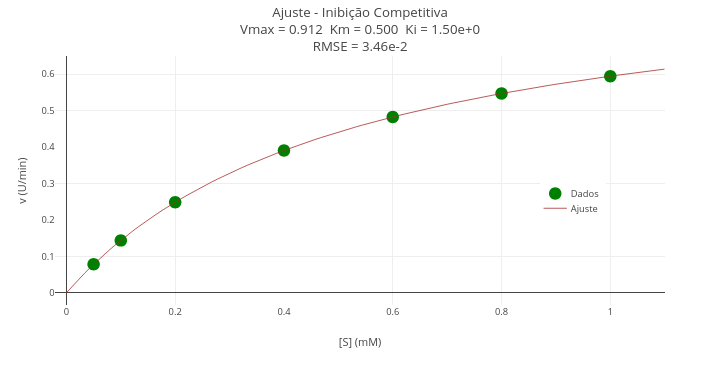
Context - Competitive inhibition in enzyme kinetics (Gauss-Newton algorithm)
Equation:
\[ \theta_{k+1} = \theta_k + \left(J^T J\right)^{-1} J^T \cdot r \]
Instructions
- Notice that the code has a true/false flag in the constant ajuste. Leave it false for representing the data only (vectors x and y, or S and v);
- In the code’s Plot area, set the constant trace so that it uses the data vectors as well;
- Click add plot to view the experimental points;
- To overlay the fitted curve, update the constants ajuste and trace (vectors S_fit and v_fit), and choose lines in mode.
4 Applications
4.1 prot.Id - Simulated experimentation for characterization of amino acids and chains
Context:
Simulator instructions
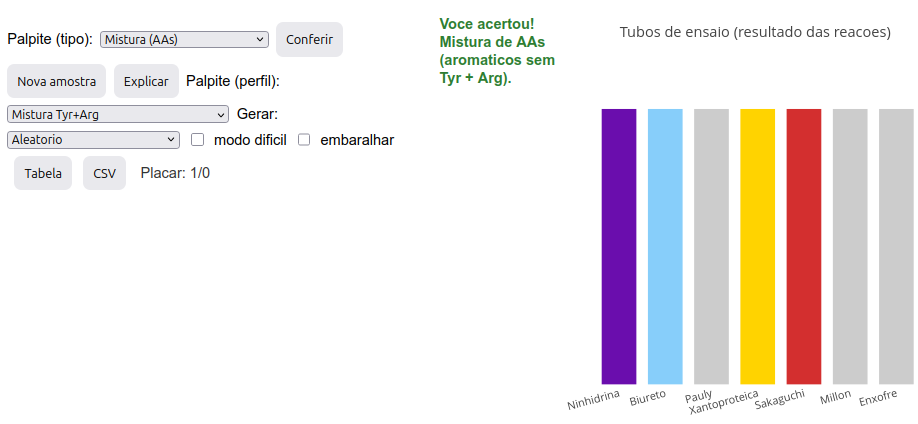
Option A: discovering the nature of a sample:
Click the image below and press “add”. A set of 7 colored bars will be generated, representing the colors obtained in test tubes for 7 distinct reactions used for protein characterization: ninhydrin, biuret, Pauly, xanthoproteic, Sakaguchi, Millon, and sulfur (Cys);
Determine the sample type based on the color pattern presented, making a guess. The guess must be by type (in the “Guess (type)” menu—AA, chain, or AA mixture) and by profile (in the “Guess (profile)” menu—random, specific AA(s) or specific mixture);
After selecting both menus, click “Check”. If correct, a “Scoreboard” right below will count the number of correct guesses/number of samples “played”;
If you miss, try again or look for the explanation of the color pattern in “Explain”;
There are two other possibilities: “Hard mode”, which hides the reaction names while keeping their sequence, and “Shuffle”, which changes the order of the tubes;
For a new sample, just click “New sample”;
Option B: discovering the reaction color pattern for a selected sample:
- Select a sample in “Generate”, and click “New sample”. A set of tubes will be generated with the expected reaction pattern for that sample.
Suggestion:
1. Try clicking "New sample" to see how often possibilities occur;
2. Try both operating options of the "game" (yes, it is a game...after all, it has a scoreboard!);
3. Try "hard mode" (without reaction labels in sequence) or "shuffle" (permutes the reaction order). Obviously, however, avoid using both at the same time!
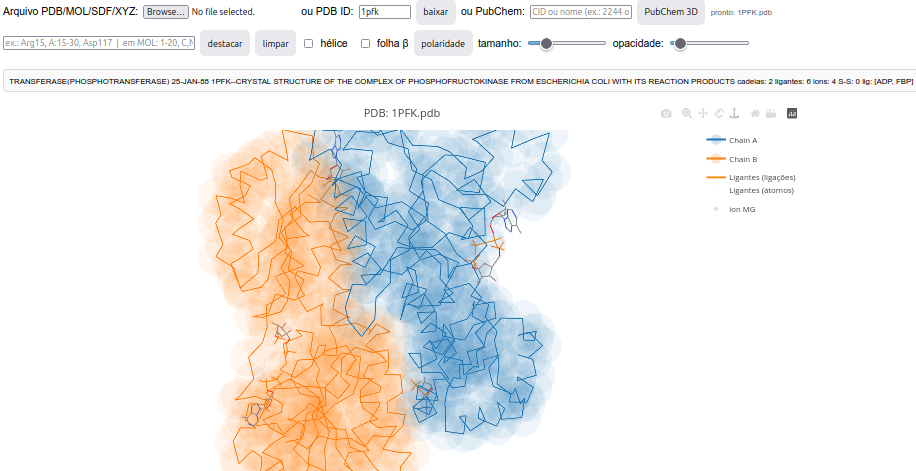
4.2 ZeMol - 3D viewer for molecular models
Context:
- models loaded online from the PDB or PubChem sites;
- models loaded from the device’s physical storage (PDB, MOL, SDF, XYZ, TXT formats);
- identification information for the PDB file (or PubChem file name);
- header containing counts of chains, ligands, ions, and disulfide bonds;
- identification of only the \(\alpha\) carbons of each residue in the polypeptide backbone, facilitating model visualization;
- automatic visualization of disulfide bonds;
- identification of residues, ligands, and ions by mouse hover over the model;
- checkbox to show protein secondary structure (helix, \(\beta\)-sheet);
- checkbox for residue polarity index;
- sliders for CA carbon size (PDB) or atoms in general, and model opacity (useful to highlight clefts, secondary structure, carbon chain, ligands, ions);
- hide/show distinct parts of the model by clicking its legend (chain, ions, helices/sheets, ligands);
- highlighting amino acids (single, range, all of one type, predefined groups in the script);
- self-contained HTML when saving, “freezing” the model in the characteristics you want to present;
- Zoom on mobile devices (one finger fixed on the screen and another dragging the model) and Pan (moving the model with two fingers on the screen)—actions also present in the icons above the model;
- optional adjustments in the script itself (sizes, colors, thicknesses, AA groups, line type and thickness, for example).
Suggestion:
1. Load a model with a PDB code, or a molecule from the PubChem site;
2. If it is a protein, try the separate and combined effects of the sliders for CA size and opacity;
3. Observe residue polarity in the polarity checkbox;
4. Try zooming in/out or moving it on the screen with the mouse, or doing it on a smartphone as instructed above;
5. Show or hide each chain or all chains, ligands, and ions by clicking the corresponding legend terms;
6. Select residues of interest in the model, such as a catalytic site, coenzyme-binding site, or regulatory residues. To do this, type in the corresponding field and click "highlight". Some suggestions:
a. Single: Arg15, ASP117, 15, A:15, A:ARG15;
b. Ranges: 15-30, A:15-30, Arg15-Asp30;
c. Multiple separated by commas: "Arg15, Asp117";
d. Groups: "aromatico", "polar", "apolar", "basico", "acido", "small".
4.3 carbo.Id - Simulated experimentation for carbohydrate characterization
Context:
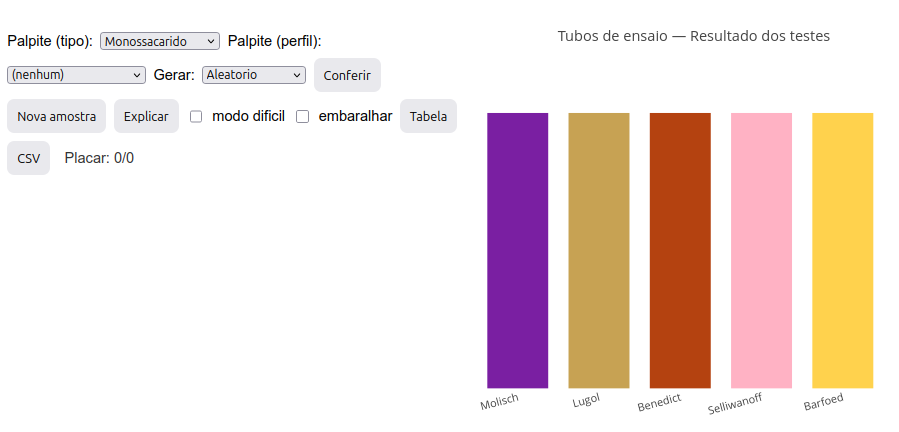
Molisch test – detects carbohydrates (purple ring); reaction with \(\alpha\)-naphthol producing furfural or hydroxymethylfurfural;
Lugol test (I\(_{2}\)/KI) – detects polysaccharides (dark blue tube);
Benedict test – detects reducing monosaccharides (and also lactose); tube with dark orange/brick precipitate;
Seliwanoff test (resorcinol) – differentiates ketoses (red tube) from aldoses (faint pink tube);
Barfoed test – differentiates mono (tube with red precipitate – cuprous oxide) from oligosaccharides (yellowish tube).
Simulator instructions
Option A: discovering the nature of a sample:
Click the image below and press “add”. A set of 5 colored bars will be generated, representing the colors obtained in test tubes for the 5 distinct reactions used for carbohydrate characterization: Molisch, Lugol, Benedict, Seliwanoff, and Barfoed.
Determine the sample type based on the tube color pattern presented, making a guess. The guess must be by type (in the “Guess (type)” menu—monosaccharide, di-oligosaccharide, polysaccharide, water) and by profile (in the “Guess (profile)” menu—reducing, non-reducing, aldose, ketose, monosaccharide, di-oligosaccharide, and polysaccharide);
After selecting both menus, click “Check”. If correct, a “Scoreboard” right below will count the number of correct guesses/number of samples “played”;
If you miss, try again or look for the explanation of the color pattern in “Explain”;
There are two other possibilities: “Hard mode”, which hides the reaction names while keeping their sequence, and “Shuffle”, which changes the order of the tubes;
For a new sample, just click “New sample”;
Option B: discovering the reaction color pattern for a selected sample:
- Select a sample in “Generate”, and click “New sample”. A set of tubes will be generated with the expected reaction pattern for that sample.
Suggestion:
1. Try clicking "New sample" to see how often possibilities occur;
2. Try both operating options of the "simulator/app/game");
3. Try "hard mode" (without reaction labels in sequence) or "shuffle" (permutes the reaction order). Obviously, however, avoid using both at the same time!
4.4 FlowForces
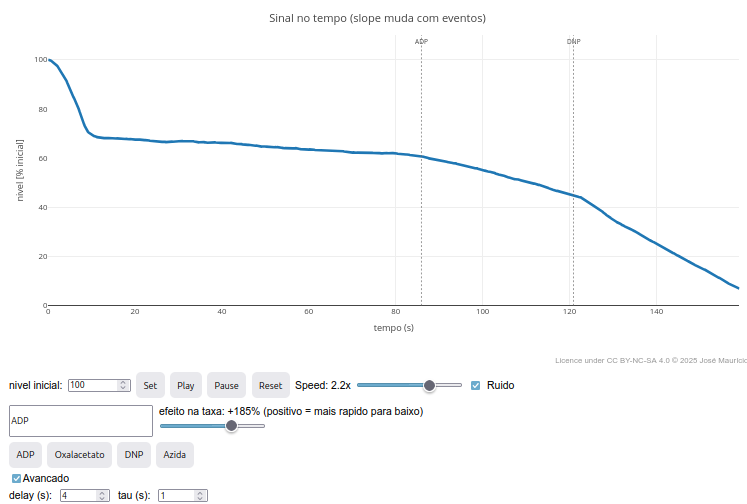
Context:
Suggestion:
1. Try changing one or more compounds with biochemical justification, for example for a metabolic pathway;
2. Test variations of "delay" for the appearance of a modifier effect by clicking the "Advanced" checkbox;
3. Try increasing the noise in "const noise", raising it 100x:
"const noise = noiseOn ? (Math.random()-0.5)*20"
4.5 MentMapa - editor for mind map and diagrams
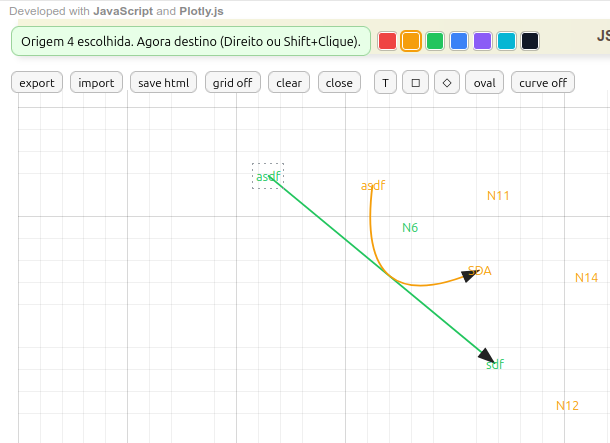
Instructions for MentMap
- Creating terms. Click with the left mouse button and type a term; press Esc to exit;
- Deleting terms. Click Ctrl+right mouse button;
- Frame on terms. Click on a frame on the bar (square, diamond, oval);
- Connecting terms by arrow. Right-click the 1st term, followed by right-click on the 2nd term;
- Curved arrow. Click curve on. Then, right-click the 1st term, followed by left-click on a point between the two terms, and finish with right-click on the 2nd term. To adjust the arrow curvature, click the little ball and drag it to touch a term or an arrow;
- Drag terms and arrows. Just click and drag.
- Drag a group of terms/arrows. Press Shift and draw the area you want to encompass the group with the left mouse button;
- Colors for terms and arrows. Select one of the 7 colors available on the top bar, and click an existing term, or a new one. Obs: the arrow color follows that of the terms;
- Grid for more precise positioning. Just click grid on (it is not saved in the final diagram);
- Saving the map (save html). Different from the usual html button, storing the interactive map is done by its own html button, on the icon bar;
- Importing maps (import). The application allows importing an already ready map in json format to continue its editing;
- Exporting maps (export). The application also allows exporting the map in json format for future study or simple keeping.
Suggestion:
1. The editor allows creating mind maps, diagrams, flowcharts, and other representations...so now the suggestion is yours !!
4.6 Gizz - Digital chalkboard
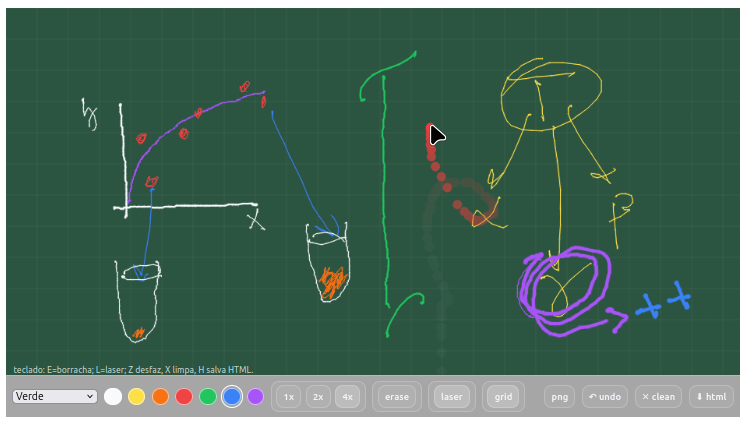
Context:
- It can be exported as a self-contained HTML file via a button on the board itself, allowing sharing of the generated image and continued editing;
- It allows customization of the code that produces it, making it possible to insert/change pens, thicknesses, and different actions not provided in the source code.
- Writing with 7 colors and 3 different thicknesses;
- Accessing 4 background planes with different coloration;
- Overlaying the background with a grid;
- Using an eraser (erase);
- Using a laser pointer to transiently locate what you want to point out;
- Saving the image as PNG, indicating date and time;
- Commands for undo (undo) and cleaning (clean) the board;
- Keyboard shortcuts: E=eraser; L=laser; Z undo, X clear, H save HTML
Suggestion:
1. You can change pen thickness in "var baseWidth = 1;" ;
2. The 1x, 2x, 4x buttons are multipliers; edit the button creation lines to change label/multiplier (e.g., "addSize('3x', 3, true)").
5 Toolkits
Important notes:
The Toolkits contain internal code for styles and panel assembly that can interfere with the Bootstrap of the web page builder used for Bioquanti (the Quarto package for R). In this sense, the clickable images for each panel point to the individual scripts for loading each Toolkit into JSPlotly.
Each panel is displayed below the JSPlotly plotting screen. This is necessary to maximize the area available for displaying the panels.
Individually, the toolkits include:
- EK-Toolkit - modeling in enzyme kinetics and inhibition;
- LeadHunt - lead-compound prospecting via enzyme inhibition;
- LB-Toolkit - analysis of ligand–protein interaction;
- GH-Toolkit - thermodynamic stability of biopolymers;
- mTherm-Toolkit - stability under perturbing conditions;
- ThermStab-Toolkit - enzyme thermostability;
- Biocel-Toolkit - microbial fuel cell performance
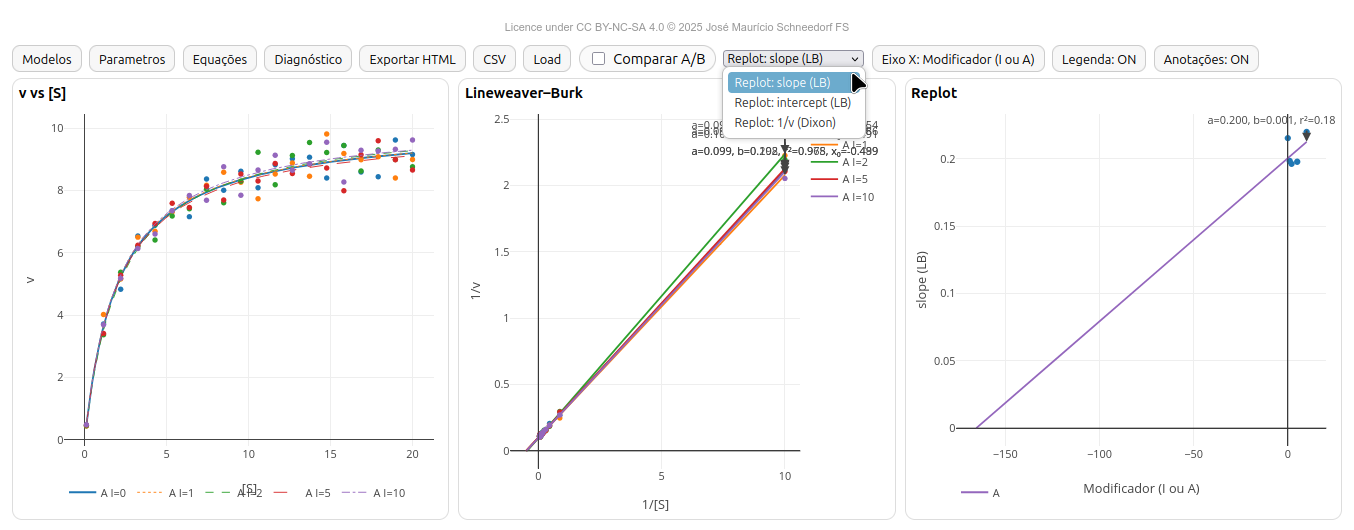
5.1 EK-Toolkit
General instructions
- Michaelis–Menten: v × [S];
- Lineweaver–Burk (LB) — 1/v vs 1/[S];
- Replot - LB slope or intercept, or Dixon ordinate (1/v)
Features
- Models: Michaelis–Menten; Substrate excess; Irreversible inhibition; Competitive inhibition; Non-competitive inhibition (pure); Mixed inhibition; Uncompetitive inhibition; Bi-substrate (Random Sequential, Ordered Sequential, Ping-Pong); Hill (cooperativity); Enzyme activation (Vmax-up); User (free equation);
- Synchronized plots: v×[S], LB (1/v vs 1/[S]) and Replot (slope, intercept or Dixon 1/v vs modifier);
- A/B comparison with consistent color palettes across plots;
- Panels: models, equations, parameters (range of [S], number of simulated points, error, modifiers, parameters;
- Configurable Replot with editable X-axis label.
- Fits: weighted linear fit in LB and replots - WLS (weighted least squares, weight: 1/y²); OLS (ordinary least squares) in Dixon.
- Ki estimation in Dixon; vertical marker at (x = -K_i).
- HTML export with interactive plots and equations; PT-BR CSV (‘;’ separator, ‘,’ decimal).
- CSV load to reconstruct plots and diagnostics; ‘Clear CSV’ to return to simulation.
- Draggable modals; clickable/editable annotations; global legend ON/OFF.
Models
Michaelis–Menten \[ v = \dfrac{V_{max} S}{K_m + S} \]
Substrate excess \[ v = \dfrac{V_{max} S}{K_m + S + \dfrac{S^2}{K_{is}}} \]
Irreversible inhibition \[ v = \dfrac{V_{max}(1-f) S}{K_m + S} \]
Competitive \[ v = \dfrac{V_{max} S}{K_m \left(1 + \dfrac{I}{K_i}\right) + S} \]
Non-competitive (pure) \[ v = \dfrac{V_{max}}{1 + \dfrac{I}{K_i}} \cdot \dfrac{S}{K_m + S} \]
Mixed \[ v = \dfrac{V_{max}}{1 + \dfrac{I}{K_{i2}}} \cdot \dfrac{S}{K_m \left(1 + \dfrac{I}{K_{i1}}\right) + S} \]
Uncompetitive \[ v = \dfrac{V_{max} S}{K_m + S\left(1 + \dfrac{I}{K_i}\right)} \]
BiBi Random Sequential \[ v = \dfrac{V_{max} A B}{K_a K_b + K_a B + K_b A + A B} \]
BiBi Ordered Sequential \[ v = \dfrac{V_{max} A B}{K_a K_b + K_b A + A B} \]
BiBi Ping-Pong \[ v = \dfrac{V_{max} A B}{K_b A + K_a B + A B} \]
Hill (cooperativity) \[ v = \dfrac{V_{max} S^n}{K^n + S^n} \]
Activator (Vmax-up) \[ v = \dfrac{V_{max}\left(1 + \dfrac{A}{K_{aA}}\right) S}{K_m + S} \]
Note:
- a (intercept): value of 1/v when 1/[S] = 0.
- b (slope): LB line slope = (K_m/V_{max}).
- K_i: inhibition constant (Dixon: (x = -K_i)).
Suggestion
1. Try comparing the pure non-competitive model to the mixed model; for this, simply set the inhibition constants equal (pure) or not equal (mixed);
2. Try a user-introduced model ("user"). To do this, select Models --> User, enter the equation, the parameters and their values. Example (partial uncompetitive activation):
Equation: (alfa*Vmax*S)/(Km+S*(1+I/Ki))
Parameters: alfa;Km;Vmax;Ki
Values: 1.7;2;10;5Models → User, enter Equation (JS), Parameters (names;) and Values (;).
- Marangoni, Alejandro G. Enzyme kinetics: a modern approach. John Wiley & Sons, 2003.
- Purich, Daniel L. Enzyme kinetics: catalysis and control: a reference of theory and best-practice methods. Elsevier, 2010.
- Leone, Francisco A. Fundamentos de cinética enzimática. Editora Appris, 2021.
- Segel, Irwin H. Enzyme kinetics: behavior and analysis of rapid equilibrium and steady state enzyme systems. Vol. 115. New York: Wiley, 1975.
5.2 LeadHunt-Toolkit
General instructions
- At the top of the app there are 4 buttons:
- reset - renews the simulation with new data and plots;
- html - saves the plots and data as an interactive file;
- diagnostic - opens a popup window with inhibition model diagnosis and parameter values;
- parameters - opens a window for simulation based on user parameters.
- There are two study modes for LeadHunt: entering kinetic and thermodynamic parameters to observe the outputs in graphs (parameters button), or characterizing parameters from the graphs (diagnostic button).
Parameters
- Ki, inhibitor dissociation equilibrium constant (desirable < 10\(\mu\)M);
- Vmax, limiting enzyme reaction rate;
- Km, Michaelis–Menten constant;
- \(\Delta\)H, enthalpy change upon inhibitor binding;
- \(\Delta\)S, entropy change upon inhibitor binding;
- \(\Delta\)Cp, heat capacity change upon inhibitor binding;
- MW, molecular weight;
- HAC, number of heavy atoms (all except hydrogen);
- HBD, number of hydrogen bond donors;
- HBA, number of hydrogen bond acceptors;
- logP, logarithm of the partition coefficient
LE, ligand efficiency:
\[ LE = \frac{\Delta G}{HAC}; \]
\[
LE = 1.37 \frac{pIC_{50}}{HAC}
\]
Where pIC\(*{50}\) = -log(IC\(*{50}\));
- LLE, lipophilic ligand efficiency (values >5 are considered good),
\[ LLE=pIC_{50}-logP \]
- BEI, binding efficiency index,
\[ BEI=\frac{pIC_{50}}{MW} \]
Where: MW, kDa; typical ranges by Lipinski’s “rule of five”: MW<=500, HBD<=5, HBA<=10, cLogP<=5. Also LE >= 0.3; LLE >= 5.
- IC\(_{50}\), inhibitor concentration giving 50% response (adequate when < 10\(\mu\)M);
- nH, Hill constant (midpoint slope; cooperativity index)
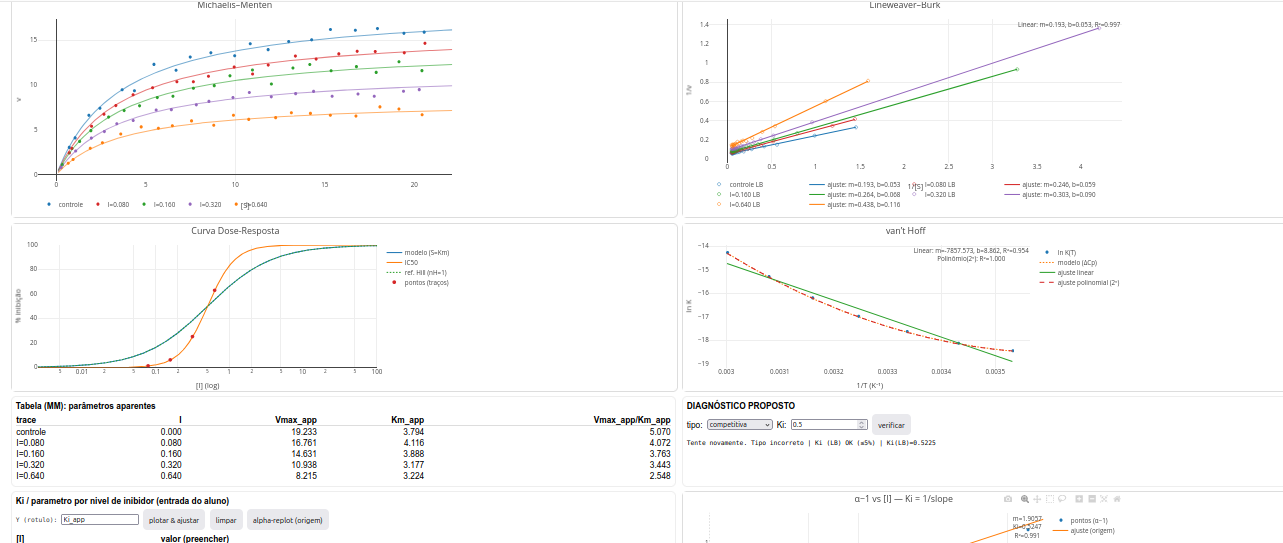
Using LeadHunt to identify a lead compound
The type of reversible inhibition (competitive, uncompetitive, pure non-competitive, mixed non-competitive);
The Ki value by constructing various secondary plots from the parameter-entry table (Ki table)—plot inhibitor concentration I versus apparent Km, Vmax, or Km/Vm. Alternatively, plot I against (\(\alpha\)-1), where \(\alpha\) represents “1+I/Ki”;
The IC\(_{50}\) value and nH (midpoint slope) from the dose–response curve (cooperativity estimate);
The \(\Delta\)H and \(\Delta\)S values from the van’t Hoff plot. If the plot is curvilinear, also the \(\Delta\)Cp value;
The LE and LLE values;
Evaluate the candidate against Lipinski’s rules.
Suggestion
1. Compare enzyme inhibition types by visual inspection of the Michaelis–Menten and Lineweaver–Burk plots, emphasizing whether "Km" and "Vmax" do or do not change between them;
2. Compute "Ki" by different methods using the field for "replot" by inhibitor level (table of "Apparent parameters"):
a) by obtaining "1/Ki" from the I x (alpha-1) plot (line slope) via the "alpha-replot" button;
b) by intercept "-Ki" from plots of I x Vmax_app, Km_app, or Km/Vm_app (depending on the inhibition model);
c) by "slope" or "intercept" from replots of Lineweaver–Burk against I
3. Try the "parameters" mode with an "nH" value different from 1 (e.g., 2.8) to evaluate dose–response curve behavior under cooperativity effects;
4. Try the "parameters" mode with a non-zero heat capacity value (e.g., -2000) to evaluate the possibility of secondary binding effects as a function of temperature, such as enzyme conformational transition;
5. Test curvature of the van’t Hoff plot for a heat-capacity change by providing positive (e.g., 2000) and negative values (e.g., -2000): if concave up, negative heat-capacity change; if concave down, positive heat-capacity change.References:
- Marangoni, Alejandro G. Enzyme kinetics: a modern approach. John Wiley & Sons, 2003.
- Copeland, Robert A. Evaluation of enzyme inhibitors in drug discovery: a guide for medicinal chemists and pharmacologists. John Wiley & Sons, 2013.
- Copeland, Robert A. Enzymes: a practical introduction to structure, mechanism, and data analysis. John Wiley & Sons, 2023.
- Leone, Francisco A. Fundamentos de cinética enzimática. Editora Appris, 2021.
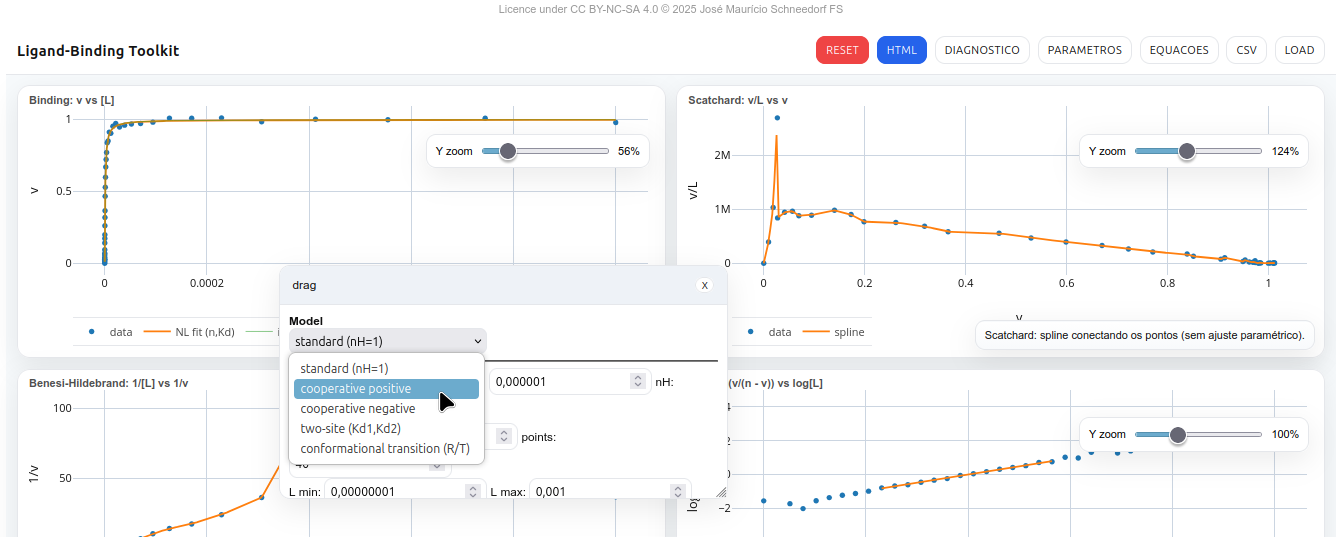
5.3 LB-Toolkit
General instructions
The LB-Toolkit is an interactive panel for analyzing ligand–receptor binding assays (protein, enzyme, nucleic acid, glycan). It allows visualization and fitting of classic binding curves (Binding, Scatchard, Benesi–Hildebrand, and Hill), and supports interactive HTML export of data and plots.
Features
L,v vectors.L,v.Adjustable parameters
n,Kd,nH,sigma error,points,Lmin,Lmax- Two-site:
n1,n2,Kd1,Kd2 - Conformational (R/T):
Kt,KdR,KdT - Toggles: draw model only, results below, Hill 10–90%, zoom sliders, autoplot
Models
Langmuir
\[
v = n,\frac{[L]}{K_d + [L]}
\]
Hill
\[ v = n,\frac{[L]^{n_H}}{K_d^{n_H} + [L]^{n_H}} \]
Linearization:
\[
\log!\left(\frac{v}{n - v}\right) = n_H \log[L] - n_H \log K_d
\]
Two-site (site heterogeneity)
\[
v = n_1,\frac{[L]}{K_{d1} + [L]} + n_2,\frac{[L]}{K_{d2} + [L]}
\]
Conformational (R/T)
\[
v = n,\frac{L/K_{dR} + K_t L/K_{dT}}{1 + L/K_{dR} + K_t(1 + L/K_{dT})}
\]
Classic linearizations
Scatchard
\[
\frac{v}{[L]} = \frac{n}{K_d} - \frac{1}{K_d} v
\]
Benesi–Hildebrand
\[ \frac{1}{v} = a,\frac{1}{[L]} + b \]
Fits and estimates
- Binding (n,Kd): non-linear fit (Gauss–Newton).
- Scatchard and Benesi: illustrative spline interpolation.
- Hill: linear regression on the 10–90% range.
5.3.0.1 Plots
- Binding: v vs [L] – theoretical curve + experimental points.
- Scatchard: v/L vs v – spline line fit to data.
- Benesi–Hildebrand: 1/[L] vs 1/v – illustrative spline.
- Hill: log(v/(n–v)) vs log[L] – linear regression with display of nH and Kd.
Suggestion
1. Use the slider (zoom bar) to enlarge/reduce the plots;
2. Try customizing the graphical representations. For example, click the "Parameters" button and change the model to negative cooperativity with two sites, assigning distinct values to them (optional).References
- Carey, Jannette. Ligand-Binding Basics: Evaluating Intermolecular Affinity, Specificity, Stoichiometry, and Cooperativity. John Wiley & Sons, 2025.
- Roque, Ana Cecília A. “Ligand-Macromolecular interactions in drug discovery.” Clifton, NJ 2010; p 572 (2010).
- Mannhold, Raimund, Hugo Kubinyi, and Gerd Folkers. Protein-ligand interactions: from molecular recognition to drug design. John Wiley & Sons, 2006.
- Klotz, Irving. Introduction to biomolecular energetics: including ligand–receptor interactions. Elsevier, 2012.
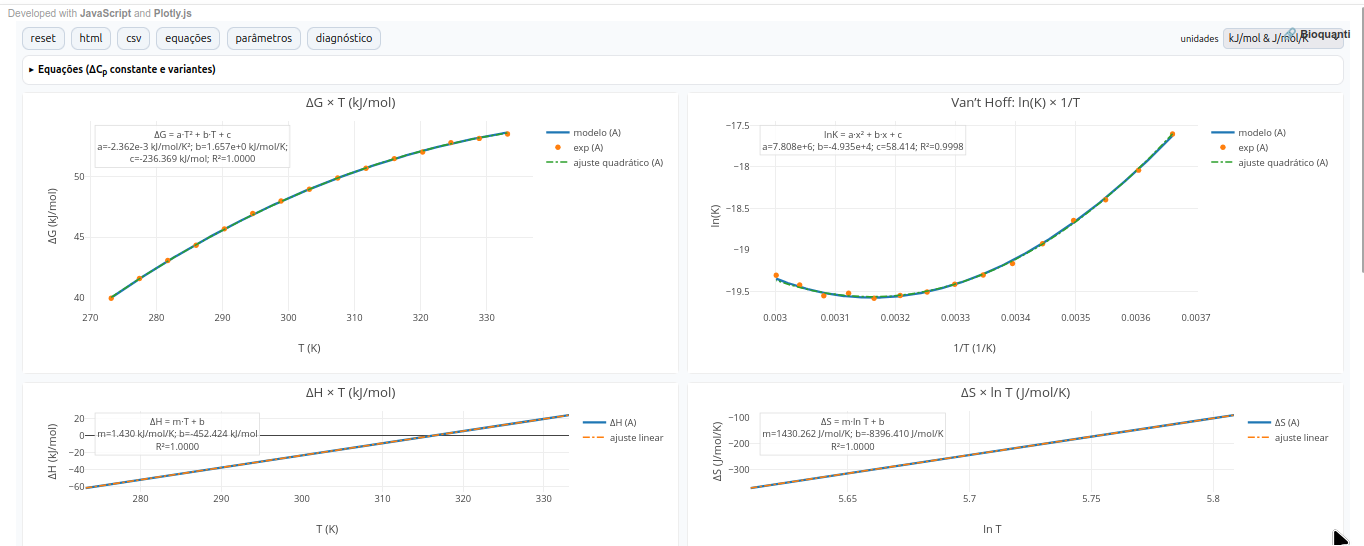
5.4 GH-Toolkit
General instructions
Parameters
1. Reference form at (T_0) (integrals with constant (ΔC_p))
Enthalpy \[ ΔH(T)=ΔH^\circ + ΔC_p,(T-T_0) \]
Entropy \[ ΔS(T)=ΔS^\circ + ΔC_p\ln!\left(\frac{T}{T_0}\right) \]
Gibbs energy (integrated Gibbs–Helmholtz) \[ ΔG(T)=ΔH^\circ - T,ΔS^\circ + ΔC_p\Big[(T-T_0)-T\ln!\left(\frac{T}{T_0}\right)\Big] \]
Equilibrium constant \[ \ln K(T)= -\frac{ΔG(T)}{R,T} \]
2. Form centered at (T_m) (stability curve via (ΔH_m) and (ΔC_p))
This form is convenient when the equilibrium temperature (T_m) (where (ΔG=0)), the enthalpy change at (T_m) ((ΔH_m)), and (ΔC_p) are known:
\[ ΔG(T)=ΔH_m!\left(1-\frac{T}{T_m}\right) + ΔC_p\Big[(T-T_m)-T\ln!\left(\frac{T}{T_m}\right)\Big]. \]
In the app, the button “generate by Eq. (T_m)” applies these relations.
3. Heat capacity with linear variation (term (ΔΔC_p))
\[ C_p(T)=C_{p,r}+ΔΔC_p,(T-T_r). \]
In the app, checking “Compare two proteins (use (ΔΔC_p)” plots protein B with (ΔC_{p,B}=ΔC_{p,A}+ΔΔC_p) and shows the consequence on the curvature of (ΔG(T)) and the shift of (T_m).
Scenarios
- Exothermic folding: (ΔH^<0); enthalpy-dominant stabilization at (T_0).
- Endothermic folding: (ΔH^>0) and (ΔS^>0) (entropy gain compensates).
- Unfolding: inverts folding signs.
- Exothermic / endothermic binding: typical settings for ligand–receptor interaction.
Suggestion
1. Note: T_m occurs when ΔG=0. In that case, a dashed vertical line is shown in the ΔG x T plot;
2. Choose a didactic "Preset" and compare its effect on stability curves:
- None: coherent random draw.
- term ↑ΔG global: makes ΔS more negative (raises ΔG globally).
- term ↓ΔC_p: flattens curvature in ΔG(T).
- psychro ↑ΔC_p: accentuates curvature.
- binder ΔC_p<0: uses negative ΔC_p (cases of reduced hydrophobic binding etc.).
3. Curvature of ΔG(T) grows with |ΔC_p|; thus, compare A vs B, and see the difference in the graphical profile ("shape") and T_m shift;
4. Note that:
- the van’t Hoff representation recovers ΔH and ΔS (when ΔC_p = 0).
- slopes/intercepts serve as consistency checks between plots
- from ΔH x T: slope gives ΔC_p.
- from ΔS x ln T: slope gives ΔC_p, and intercept is compatible with ΔS and T_0.References
- Atkins, Peter William, et al. Physical chemistry for the life sciences. Oxford University Press, 2023.
- Hammes, Gordon G. Thermodynamics and kinetics for the biological sciences. John Wiley & Sons, 2000.
- Cooper, Alan. Biophysical chemistry. No. 24. Royal Society of Chemistry, 2011.
- Creighton, T. The Biophysical Chemistry of Nucleic Acids & Proteins. Helvetian Press, 2010.
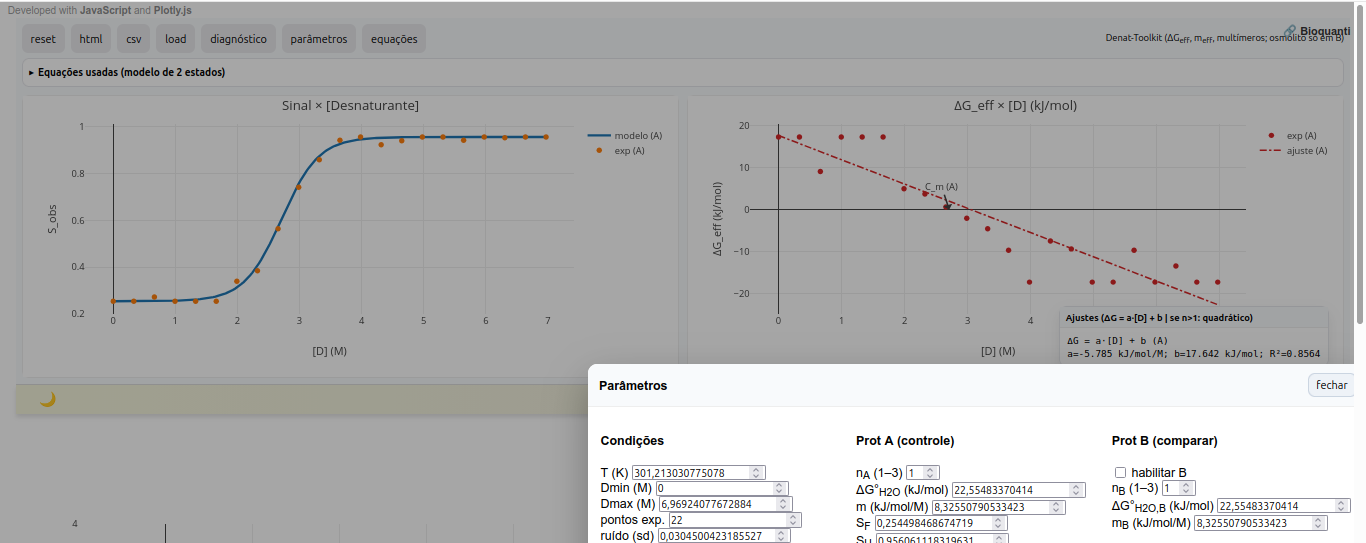
5.5 mTherm-Toolkit
General instructions
Parameters
Standard free energy vs. ([D]) — monomer (LEA)
\[ \Delta G^\circ(D)=\Delta G^\circ(\mathrm{H_2O})-m,D \tag{2.1} \]
Multimer (F_n nU) — mass action (solve (f_U))
\[
\Delta G^\circ + RT\ln!\left(\frac{n,f_U^{,n}}{P_t^{,n-1}(1-f_U)}\right)=0
\tag{2.2}
\]
- (n): number of monomers in (F_n);
- (P_t): monomer-equivalent concentration (in M);
- Eq. (2.2) is solved numerically (bisection) for (0<f_U<1) at each (D);
Effective free energy
\[ \Delta G_{\mathrm{eff}}(D)=-RT\ln!\left(\frac{f_U}{1-f_U}\right) \tag{2.3} \]
Effective slope (m_{}) (dependent on (D))
\[ m_{\mathrm{eff}}(D) = \frac{m}{,n + (1-n), f_U(D),}\quad \left(\approx \frac{m}{n}\ \text{on the native side; } \approx m \text{ on the denatured side}\right) \tag{2.4} \]
Effective intercept in water (approximation for multimers)
$$ G_{}()
n
RT!(1-1n)!P_t $$
Osmolytes
\[ m_{\mathrm{eff}} = m ,\bigl(1 + k_{\mathrm{osm}},[\mathrm{osm}]\bigr) \tag{6.1} \]
where (k_{}) is an adjustable strength parameter (typically 0.05–0.2 M\(^{-1}\)). This term shifts the curve without imposing a specific mechanism, enabling didactic exploration.
Fits and parameters
- Temperature (T) (K) and (P_t) (µM, monomer equivalent);
- Energy in water (()G\(^{H_2O}\)) and slope (m);
- Multiplicity (n): (1 n ) for A and, independently, for B;
- Baseline signals (S_F, S_U) and noise;
- A/B comparison: second protein can have different (n), (m), ()G\(^{o}\), and osmolyte settings.
Suggestion
1. Observe curvature in the ΔG x [D] plot. Values of n>1 suggest a multimeric state for the biopolymer;
2. Note that the midpoint C_m, where f_U is approximately 0.5, is marked in the diagnostic for reference;
3. Compare osmolyte effects by comparing proteins or conditions A and B, changing osmolyte concentration and its empirical constant "k_osm".References
- Park, Chiwook, and Susan Marqusee. “Analysis of the stability of multimeric proteins by effective ΔG and effective m‐values.” Protein science 13.9 (2004): 2553-2558.
- Holthauzen, Luis Marcelo F., et al. “Protein stability in the presence of cosolutes.” Methods in Enzymology. Vol. 492. Academic Press, 2011. 61-125.
- Scholtz, J. Martin, Gerald R. Grimsley, and C. Nick Pace. “Solvent denaturation of proteins and interpretations of the m value.” Methods in enzymology. Vol. 466. Academic Press, 2009. 549-565.
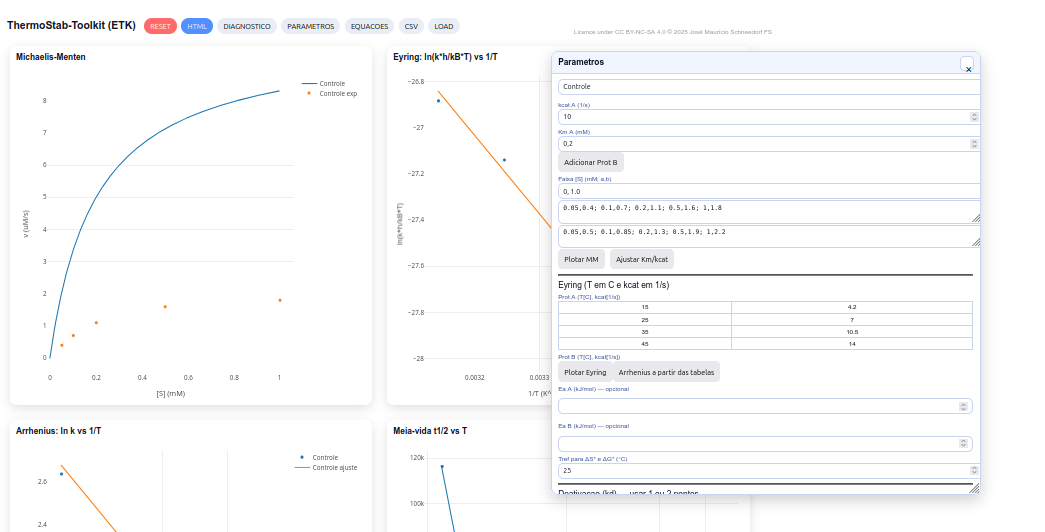
5.6 ThermStab - Toolkit
General instructions
Parameters
Michaelis–Menten (MM)
\[ v = \frac{V_{\max}[S]}{K_m + [S]} \]
\[
V_{\max} = k_{\text{cat}} \cdot E_0 \qquad (E_0 = 1.0\ \mu\text{M})
\]
\[ \Delta G^{\ddagger}_{ES} = -RT \ln \left(\frac{1}{K_m}\right) \]
\[ \Delta G^{\ddagger}*{ET} = -RT \ln \left(\frac{k*{\text{cat}}}{K_m}\right) \]
Eyring equation
\[ \ln \left(\frac{k h}{k_B T}\right) = -\frac{\Delta H^{\ddagger}}{RT} + \frac{\Delta S^{\ddagger}}{R} \]
\[
\Delta G^{\ddagger} = \Delta H^{\ddagger} - T\Delta S^{\ddagger}
\]
Arrhenius equation
\[
\ln k = -\frac{E_a}{RT} + \ln A
\]
Enzyme half-life
\[
t_{1/2} = \frac{\ln 2}{k_d}
\]
Binding equilibrium
\[
K_d = \frac{[E][S]}{[ES]}
\]
\[ \ln K_d(T) = -\frac{\Delta H^\circ}{RT} + \frac{\Delta S^\circ}{R} \]
Kinetic relations
\[
K_m \approx \frac{k_{-1} + k_{\text{cat}}}{k_1}
\]
\[
K_d \approx \frac{k_{-1}}{k_1}
\]
\[
k_{-1}(T) \approx k_1,K_d(T)
\]
Relation between stability and catalysis
\[
k_d(T) \approx k_1 K_d(T) - k_{\text{cat}}(T)
\]
\[
t_{1/2}(T) = \frac{\ln 2}{k_d(T)}
\]
Didactic approximation
\[
\ln t_{1/2} \approx \frac{\Delta H^\circ}{R}\left(\frac{1}{T}\right)* \frac{\Delta S^\circ}{R} + \text{constant}
\]
Suggestion
1. Use the RESET button and discuss the shape of the MM curve and the influence of Km and kcat;
2. Explore Eyring and Arrhenius relations to obtain ΔH‡, ΔS‡, ΔG‡ and Ea from slopes and intercepts of the plots;
3. Explore the effect of temperature and compare with free mode (t₁/₂) in class.References
- Bhatti, Haq Nawaz, et al. “Effect of aniline coupling on kinetic and thermodynamic properties of Fusarium solani glucoamylase.” Applied microbiology and biotechnology 73.6 (2007): 1290-1298.
- Lonhienne, Thierry, Charles Gerday, and Georges Feller. “Psychrophilic enzymes: revisiting the thermodynamic parameters of activation may explain local flexibility.” Biochimica et Biophysica Acta (BBA)-Protein Structure and Molecular Enzymology 1543.1 (2000): 1-10.
5.7 Biocel-Toolkit
General instructions
Parameters
Thevenin model
Current (I) and the voltage across the load resistor (V) are given by:
\[
I = \frac{E_{OCV}}{R_{int} + R_{ext}}, \qquad
V = I , R_{ext}
\]
Electrical power and maximum condition
\[
P = E_{OCV} , I - R_{int} I^2
\]
\[
P = \frac{E_{OCV}^2 , R_{ext}}{(R_{int} + R_{ext})^2}
\]
The maximum power point occurs when (R_{ext} = R_{int}), known as the Jacobi condition, yielding:
\[
P_{\text{max}} = \frac{E_{OCV}^2}{4 R_{int}}
\]
Activation and concentration overpotentials
\[ V_{pol} = V_{lin} - \eta_{act} - \eta_{conc} \]
where:
\[
\eta_{act} = A \ln!\big(1 + \frac{I}{I_0}\big), \qquad
\eta_{conc} = C , \frac{(I/I_{lim})^2}{1 - I/I_{lim}}
\]
Time dynamics
\[
E_{OCV}(t) = E_{\max}\big(1 - e^{-k_{act}t}\big) s(f)
\]
\[
R_{int}(t) = R_0 \big(1 + \beta (1 - f)\big)
\]
Where:
\[ s(f) = 0.40 + 0.60f, \quad f = \frac{Q}{Q_{theor}} \]
Charges and efficiencies
\[
Q_{theor} = n_{\text{gluc}} , n_e , F
\]
Where: (F) is Faraday’s constant and (n_e = 24) is the number of electrons per mole of glucose.
\[ \eta_Q = 100 \times \frac{\int I,dt}{Q_{theor}} \quad [%] \]
\[ \eta_E = 100 \times \frac{\int IV,dt}{Q_{theor} E_{\max}} \quad [%] \]
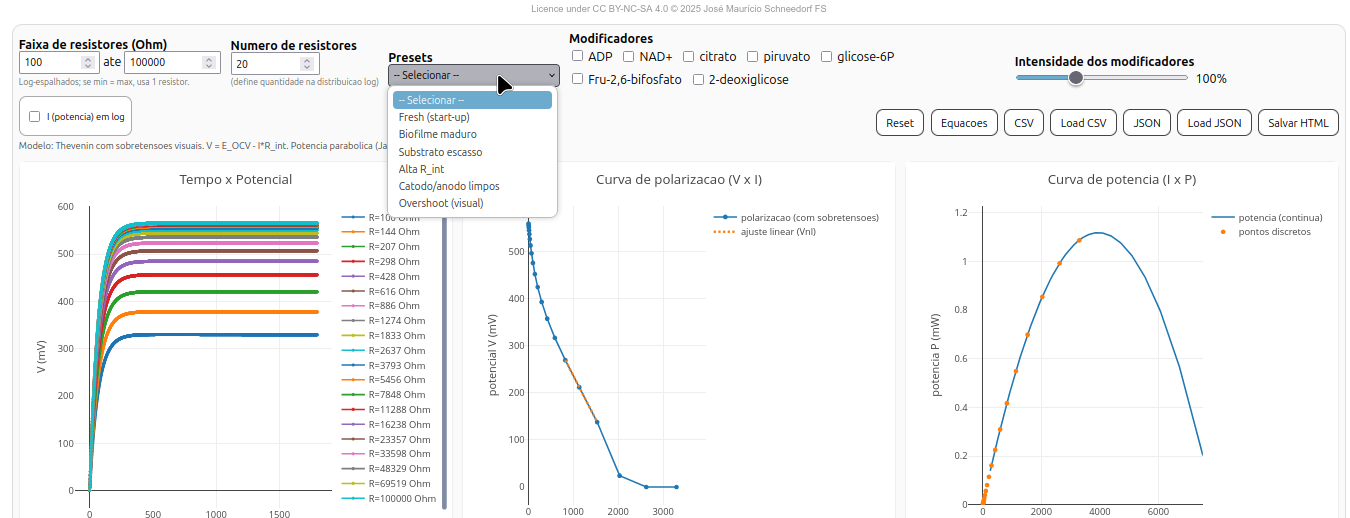
Interface and features
The BioFuelCell interface is divided into three plot panels and a parameter section:
- Time × Potential (t × V) — shows output potential evolution over time for each external resistor;
- Polarization curve (V × I) — shows voltage decay as a function of current, with or without overpotentials;
- Power curve (I × P) — computes power dissipated in the load and highlights the maximum power point.
- (R_{int,lin}) — internal resistance by linear fit of the real curve;
- (R_{int,ideal}) — theoretical internal resistance (Thevenin model);
- (P_{}) and (R_{P}) — maximum power and corresponding resistance;
- (_Q) and (_E) — charge and energy efficiencies.
5.7.1 Parameters and modifiers
- Resistor range — minimum and maximum (R_{ext}) (in Ω), distributed logarithmically;
- Modifier intensity — from 0% to 300%;
- Log scale for power — toggles current axis to log scale;
- Experimental presets — predefined simulated operating conditions.
Suggestion
1. Observe the intersection point between the polarization curve and the ideal parabola. This point indicates the balance between electron supply and consumption;
2. Note the initial slope of the V×I curve, which represents the internal resistance (R_int) of the biocell;
3. Check the system’s internal efficiency by observing the position of the peak in the power plot;
4. Try some of the experimental "Presets" (mature biofilm, substrate excess, etc.) and evaluate how conditions affect the electrochemical performance shown in plots and results.
The potential vs time plot should appear.
References
- Logan, B. E. Microbial Fuel Cells. John Wiley & Sons, 2008.
- Bard, A. J., & Faulkner, L. R. Electrochemical Methods: Fundamentals and Applications. Wiley, 2022;
- Silveira, G., & Schneedorf, J. M. “Evaluation of kefir as a new anodic biocatalyst consortium for microbial fuel cell.” Applied Biochemistry and Biotechnology, 185(4), 1118–1131 (2018).
6 Arduino & Biocell
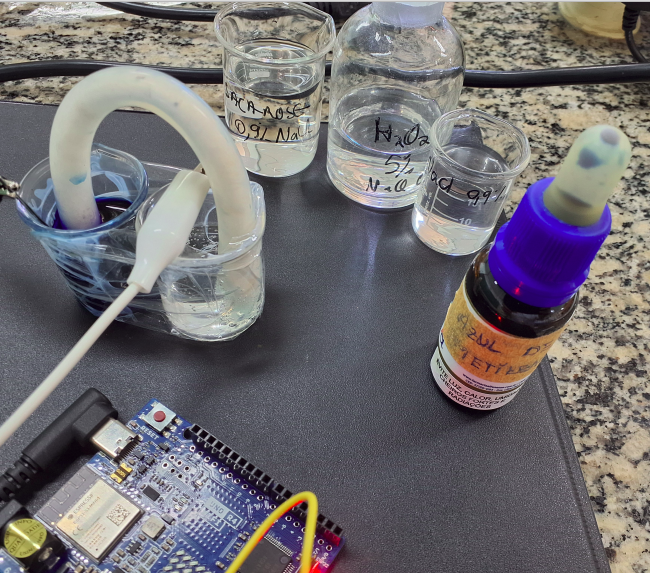

6.1 Serial cable
"** QUICK TIP ** ":
1. Connect the Arduino board;
2. Open a CMD or Terminal and type:
"python3 -m http.server" ;
3. Upload the sketch;
4. Open the software in Chrome (Firefox is problematic)
5. Connect to the Arduino port in the software6.2 Internet of Things - IoT
Plotly.js library for tablets and phones. The image below illustrates an experiment conducted with a microbial fuel cell, interfaced via Wi-Fi to an Arduino Uno R4 WiFi board.
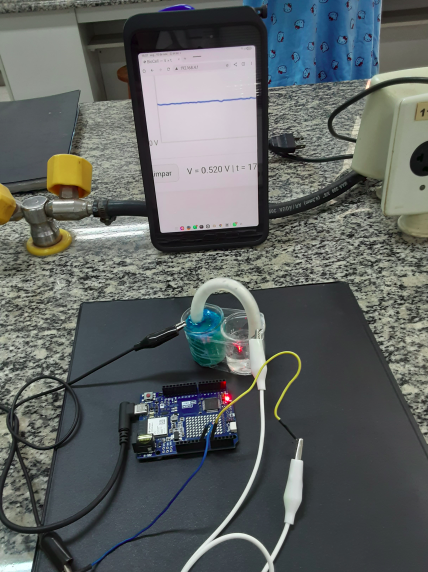

"** QUICK TIP ** ":
1. Connect the Arduino board;
2. Upload the sketch;
3. Open Serial Monitor;
4. Optional: press Reset on the Arduino for ~2s;
5. Access the specific network on the phone
In Wi-Fi, choose Biocel_no.1
Password: 12345678 ;
6. Open the link http://192.168.4.1
The potential vs time plot should appear.