# Data input
1. After selecting "AIC", the program requests the following values (INPUT):
n, kA, kB, ssA, ssB
2. After ENTER, the program computes:
"AIC_A, AIC_B, delta, prob.B"AIC – Akaike Information Criterion and Enzyme Inhibition
1 Equation:
\[ AIC = n \cdot \ln!\left(\frac{SSE}{n}\right) + 2k + \left[\frac{2k(k+1)}{n-k-1}\right] \]
Where:
- k = p + 1 (with p being the number of model parameters);
- SSE = sum of squared residuals from the fit.
\[
probab.B = \frac{e^{-0.5 \cdot \Delta AIC}}{1 + e^{-0.5 \cdot \Delta AIC}}
\]
Where:
- probab. B = probability that model B is better than model A (range 0–1);
- \(\Delta\)AIC = difference AIC\(_B\) − AIC\(_A\).
2 Files
3 Usage:
4 Example
Table 1 – Experimental enzyme inhibition data (n = 105). Source: Bezerra et al., 2013.
| Type | Control | Competitive | Pure non-competitive | Mixed non-competitive | Uncompetitive |
|---|---|---|---|---|---|
| SSE | 1540.1 | 613.1 | 1122.7 | 613.2 | 1436.4 |
| p | 2 | 3 | 3 | 4 | 3 |

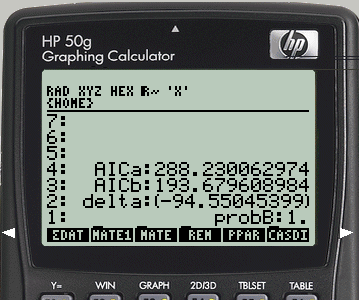
Table 2 — Results of model comparison using the AIC criterion.
| Model A / B | AIC\(_A\) | AIC\(_B\) | \(\Delta\) | Probab. B |
|---|---|---|---|---|
| Control / Competitive | 288.2 | 193.7 | -94.6 | 1.0 |
| Control / Mixed NComp. | 288.2 | 195.9 | -92.3 | 1.0 |
| Competitive / Mixed NComp. | 193.7 | 195.9 | 2.2 | 0.25 |
References
Akaike, Hirotugu. A new look at the statistical model identification. IEEE Transactions on Automatic Control, 19(6), 716–723, 2003.
Bezerra, Rui M. F., Irene Fraga, and Albino A. Dias. Utilization of integrated Michaelis–Menten equations for enzyme inhibition diagnosis and determination of kinetic constants using the Solver supplement of Microsoft Office Excel. Computer Methods and Programs in Biomedicine, 109(1), 26–31, 2013.